MHETase
The Enzyme MHETase is a hydrolase, which was discovered in 2016. It cleaves Mono-(2-hydroxyethyl)terephthalic acid, the PET degradation product by PETase, to ethylene glycol and terephthalic acid.[1] This pair of enzymes, PETase and MHETase, enable the bacterium Ideonella sakaiensis to live on the plastic PET as sole carbon source.
| MHETase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| EC number | 3.1.1.102 | ||||||||
| Alt. names | MHET hydrolase, monohydroxyethyl terephthalate hydrolase | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
Chemical reaction
The first enzyme of the PET degradation pathway, PETase, cleaves this plastic into the intermediates MHET (Mono-(2-hydroxyethyl)terephthalic acid) and minor amounts BHET (Bis-(2-hydroxyethyl)terephthalic acid). MHETase hydrolyses the ester bond of MHET forming terephthalic acid and ethylene glycol.
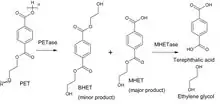
Besides its natural substrate MHET the chromogenic substrate MpNPT, Mono-p-nitrophenyl-terephthalate, is also hydrolyzed well. This can be used to measure the enzymatic activity and determine the kinetic parameters. Ferulate and gallate esters, substrates of the closest relatives in the tannase family, are not converted. p-Nitrophenyl ester of aliphatic monocarboxylic acids like the widely used esterase substrate p-nitrophenyl acetate are not hydrolyzed either.
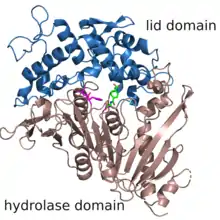
Structure
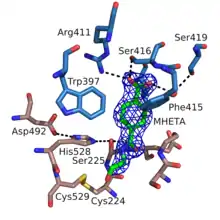
The structure of MHETase was solved in 2019.[2] It shows the common fold of the alpha/beta hydrolase superfamily. According to the classification in the ESTHER database, MHETase belongs to the family of tannases within block X.[3] This family mainly contains tannases und feruloyl esterases. The enzyme consists of two domains: the hydrolase domain harbors the catalytic residues Ser225, His528 and Asp492; the lid domain contributes most of the residues of the substrate binding site.
External links
References
- Yoshida S, Hiraga K, Takehana T, Taniguchi I, Yamaji H, Maeda Y, et al. (March 2016). "A bacterium that degrades and assimilates poly(ethylene terephthalate)". Science. 351 (6278): 1196–9. doi:10.1126/science.aad6359. PMID 26965627.
- Palm GJ, Reisky L, Böttcher D, Müller H, Michels EA, Walczak MC, et al. (April 2019). "Structure of the plastic-degrading Ideonella sakaiensis MHETase bound to a substrate". Nature Communications. 10 (1): 1717. doi:10.1038/s41467-019-09326-3. PMC 6461665. PMID 30979881.
- Renault L, Nègre V, Hotelier T, Cousin X, Marchot P, Chatonnet A (December 2005). "New friendly tools for users of ESTHER, the database of the alpha/beta-hydrolase fold superfamily of proteins". Chemico-Biological Interactions. 157-158: 339–43. doi:10.1016/j.cbi.2005.10.100. PMID 16297901.