Paralytic (gene)
Paralytic is a gene in the fruit fly, Drosophila melanogaster, which encodes a voltage gated sodium channel within D. melanogaster neurons.[1] This gene is essential for locomotive activity in the fly.[1] There are 9 different para alleles, composed of a minimum of 26 exons within over 78kb[2] of genomic DNA.[3] The para gene undergoes alternative splicing to produce subtypes of the channel protein.[3] Flies with mutant forms of paralytic are used in fly models of seizures, since seizures can be easily induced in these flies.[4]
| Paralytic | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | para | ||||||
| Alt. symbols | bss, sbl, olfD, DmNav, DmNav1, DmNav | ||||||
| NCBI gene | 32619 | ||||||
| UniProt | P35500 | ||||||
| Other data | |||||||
| Locus | Chr. X 16,455,230 - 16,533,368 | ||||||
| |||||||
Gene
The para gene is located on the X chromosome within the Drosophila genome.[5] There are 26 para exons, 13 are constitutively expressed in the transcript, while 15 are alternatively spliced.[6] Alternative splicing allows for the formation of 60 unique transcripts and 57 unique polypeptides.[6] The independent splicing of 11 exons allows for the unique cytoplasmic loops, the alternative splicing also can effect the Na+ channel kinetics,[6] such as the varying gating conductivities.[1] The mature mRNA transcript only includes one of C or D exonic region and only one of K or L exonic region, as they code for the same or similar regions.[6] Neurons containing para exon L, show an increase in firing frequency which is associated with increase seizure susceptibility.[6] Channel kinetics are influenced by splicing, that not only changes protein structure but can allow for varying modifications, like differential binding of cofactors.[7]
Known Mutant Variants
Currently there are 117 known allele variants within the para locus,[6] a few are mentioned below.
| Mutant | Properties | Mutation | Phenotype |
|---|---|---|---|
| parabss1 | gain-of-function | mis-sense substitution in S3 of HD4 | leads to seizures |
| paraGEFS+ | sustained depolarization in GABA neurons | K→T knock-in at S2 of HD2 | temperature susceptibility for seizures |
| paraDS | reduction in Na+ current | S→R knock-in at S1 of HD2 | temperature susceptibility for seizures |
| paraJS | reduction of transcription | transposon insertion 3' UTR | seizure-suppressor |
Protein
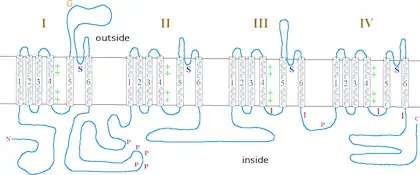
This proteins forms a sodium-selective ion channel, that relies on an electrochemical gradient.[1] The protein consists of four homology domains, HD1, HD2, HD3 and HD4. Each homology domain has six alpha helical segments, S1-S6.[6] The small alpha-helical region between S5 and S6 is known as the channel pore. Mutations within this region may be responsible for ion selectivity.[6] The cytoplasmic loop between HD3 and HD4 is responsible for fast inactivation and blocking conductance.[6] The voltage sensor 4 in HD III is partially formed by exon L and K.[7] The alternative splicing at this locus causes a difference in the charged current at this channel.[7] Exon L produces 8% of the transient current, which falls to 2% when the K exon is incorporated.[7]
Function
Paralytic encodes a protein channel which transfers sodium ions into neurons and is activated in response to changes in the voltage across a membrane[1] to propagate an action potential.[3] The paralytic protein has been found in the thoracic-abdominal ganglion, eye tissues and cortical regions in the brain.[1]
Role in seizure models
Flies with certain mutations in para gene are used as models for studying seizures and epilepsy, as they are much more prone to seizures than regular flies.[4] Some of these mutant para genotypes are cause either severe sensitivity to seizures, or act as seizure suppressors.[6] In these mutant flies, seizures can be induced by mechanical shock, electrical shock, or high-frequency visual stimuli such as strobe lights.
A number of mutations in paralytic have been described which can cause this increased sensitivity to seizures. Some of these, such as bss1 and bss2 can be caused by a single point mutation in the paralytic gene which makes the channel less able to inactivate itself after being activated.[4]
Understanding the genetic and environmental influences on the seizures in mutant para flies, has proved to be a trackable system in understanding the complexity in human seizure models.[6]
Evolution
Voltage-gated sodium channels are highly conserved across lineages. The exons in specific, are conserved across many diverged groups of species, this seems to indicate physiological importance.[7] Insect species have only one a single sodium channel gene which encodes the mammalian equivalent of α subunit. Insects like D. melanogaster take advantage of alternative splicing and RNA editing to generate distinct gating properties of sodium channels.[8]
Human orthologs
The most closely related genes to paralytic in humans are SCN1A, SCN8A and SCN2A, all of which are genes that encode sodium channels.[1][6] Mutations in the human orthologs have been linked to seizure disorders and cognitive defects.[9] Fly models can be used to study branches of human epilepsy, by using GEFS+ mutations at SCN1A gene for knock-in's at the para locus in D. melanogaster.[6]
See also
References
- "Dmel\para". FlyBase. Retrieved 28 March 2017.
- "D. melanogaster chrX:16,455,230-16,533,368 - UCSC Genome Browser v357". genome.ucsc.edu. Retrieved 2017-11-22.
- Loughney K, Kreber R, Ganetzky B (September 1989). "Molecular analysis of the para locus, a sodium channel gene in Drosophila". Cell. 58 (6): 1143–54. doi:10.1016/0092-8674(89)90512-6. PMID 2550145.
- Parker L, Padilla M, Du Y, Dong K, Tanouye MA (February 2011). "Drosophila as a model for epilepsy: bss is a gain-of-function mutation in the para sodium channel gene that leads to seizures". Genetics. 187 (2): 523–34. doi:10.1534/genetics.110.123299. PMC 3030494. PMID 21115970.
- "para paralytic [Drosophila melanogaster (fruit fly)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-11-22.
- Kroll JR, Saras A, Tanouye MA (December 2015). "Drosophila sodium channel mutations: Contributions to seizure-susceptibility". Experimental Neurology. 274 (Pt A): 80–7. doi:10.1016/j.expneurol.2015.06.018. PMC 4644469. PMID 26093037.
- Lin WH, Wright DE, Muraro NI, Baines RA (September 2009). "Alternative splicing in the voltage-gated sodium channel DmNav regulates activation, inactivation, and persistent current". Journal of Neurophysiology. 102 (3): 1994–2006. doi:10.1152/jn.00613.2009. PMC 2746785. PMID 19625535.
- "Paralytic". www.sdbonline.org. Retrieved 2017-11-22.
- Veeramah KR, O'Brien JE, Meisler MH, Cheng X, Dib-Hajj SD, Waxman SG, Talwar D, Girirajan S, Eichler EE, Restifo LL, Erickson RP, Hammer MF (March 2012). "De novo pathogenic SCN8A mutation identified by whole-genome sequencing of a family quartet affected by infantile epileptic encephalopathy and SUDEP". American Journal of Human Genetics. 90 (3): 502–10. doi:10.1016/j.ajhg.2012.01.006. PMC 3309181. PMID 22365152.