Ribonuclease V1
Ribonuclease V1 (RNase V1) is a ribonuclease enzyme found in the venom of the Caspian cobra (Naja oxiana).[1] It cleaves double-stranded RNA in a non-sequence-specific manner, usually requiring a substrate of at least six stacked nucleotides.[2] Like many ribonucleases, the enzyme requires the presence of magnesium ions for activity.[3]
Laboratory use
Purified RNase V1 is a commonly used reagent in molecular biology experiments. In conjunction with other ribonucleases that cleave single-stranded RNA after specific nucleotides or sequences – such as RNase T1 and RNase I – it can be used to map internal interactions in large RNA molecules with complex secondary structure or to perform footprinting experiments on macromolecular complexes containing RNA.[3]
RNase V1 is the only commonly used laboratory RNase that provides positive evidence for the presence of double-stranded helical conformations in target RNA.[4] Because RNase V1 has some activity against RNA that is base-paired but single-stranded,[5] dual susceptibility to both RNase V1 and RNase I at a single site in a target RNA molecule provides evidence of this relatively unusual conformation found in RNA loops.[6]
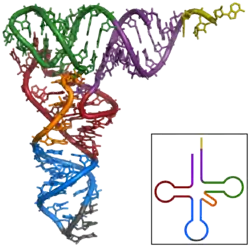
Structural discoveries
RNase V1 played a particularly important role in the elucidation of the distinctive stem-loop structure of transfer RNA.[1][7] It has also been extensively used to study the highly structured RNA genomes of retroviruses, such as hepatitis C,[8] dengue virus,[9] and HIV.[10] Together with S1 nuclease, which specifically cleaves single-stranded RNA, it can be used to profile the secondary structure propensities of messenger RNA molecules, a procedure that can be applied to whole transcriptomes when paired with deep sequencing.[11][12]
References
- Favorova OO, Fasiolo F, Keith G, Vassilenko SK, Ebel JP (February 1981). "Partial digestion of tRNA--aminoacyl-tRNA synthetase complexes with cobra venom ribonuclease". Biochemistry. 20 (4): 1006–11. doi:10.1021/bi00507a055. PMID 7011369.
- Ying, Shao Yao, ed. (2006-01-01). MicroRNA Protocols. Humana Press. p. 23. ISBN 9781597451239.
- Nilsen TW (April 2013). "RNA structure determination using nuclease digestion". Cold Spring Harbor Protocols. 2013 (4): 379–82. doi:10.1101/pdb.prot072330. PMID 23547152.
- Duval, Melodie; Romilly, Cedric; Helfer, Anne-Catherine; Fuchsbauer, Olivier; Romby, Pascale; Marzi, Stefano (2013). Klostermeier, Dagmar; Hammann, Christian (eds.). RNA Structure and Folding: Biophysical Techniques and Prediction Methods. Walter de Gruyter. p. 32. ISBN 9783110284959.
- Lowman HB, Draper DE (April 1986). "On the recognition of helical RNA by cobra venom V1 nuclease". The Journal of Biological Chemistry. 261 (12): 5396–403. PMID 2420800.
- Chaulk SG, Xu Z, Glover MJ, Fahlman RP (April 2014). "MicroRNA miR-92a-1 biogenesis and mRNA targeting is modulated by a tertiary contact within the miR-17~92 microRNA cluster". Nucleic Acids Research. 42 (8): 5234–44. doi:10.1093/nar/gku133. PMC 4005684. PMID 24520115.
- Lockard RE, Kumar A (October 1981). "Mapping tRNA structure in solution using double-strand-specific ribonuclease V1 from cobra venom". Nucleic Acids Research. 9 (19): 5125–40. doi:10.1093/nar/9.19.5125. PMC 327503. PMID 7031604.
- Blight KJ, Rice CM (October 1997). "Secondary structure determination of the conserved 98-base sequence at the 3' terminus of hepatitis C virus genome RNA". Journal of Virology. 71 (10): 7345–52. PMC 192079. PMID 9311812.
- Polacek C, Foley JE, Harris E (January 2009). "Conformational changes in the solution structure of the dengue virus 5' end in the presence and absence of the 3' untranslated region". Journal of Virology. 83 (2): 1161–6. doi:10.1128/JVI.01362-08. PMC 2612390. PMID 19004957.
- Harrison GP, Lever AM (July 1992). "The human immunodeficiency virus type 1 packaging signal and major splice donor region have a conserved stable secondary structure". Journal of Virology. 66 (7): 4144–53. PMC 241217. PMID 1602537..
- Kertesz M, Wan Y, Mazor E, Rinn JL, Nutter RC, Chang HY, Segal E (September 2010). "Genome-wide measurement of RNA secondary structure in yeast". Nature. 467 (7311): 103–7. doi:10.1038/nature09322. PMC 3847670. PMID 20811459.
- Silverman, Ian M.; Berkowitz, Nathan D.; Gosai, Sager J.; Gregory, Brian D. (2016). "Genome-Wide Approaches for RNA Structure Probing". In Yeo, Gene W. (ed.). RNA Processing. Springer. pp. 29–59. ISBN 978-3-319-29071-3.
.jpg.webp)