Five prime untranslated region
The 5′ untranslated region (5′ UTR) (also known as a leader sequence or leader RNA) is the region of an mRNA that is directly upstream from the initiation codon. This region is important for the regulation of translation of a transcript by differing mechanisms in viruses, prokaryotes and eukaryotes. While called untranslated, the 5′ UTR or a portion of it is sometimes translated into a protein product. This product can then regulate the translation of the main coding sequence of the mRNA. In many organisms, however, the 5′ UTR is completely untranslated, instead forming complex secondary structure to regulate translation.
| 5′ untranslated region | |
|---|---|
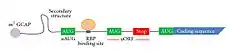 The general structure of the 5′ UTR of a transcript in eukaryotic organism (specifically humans) | |
| Identifiers | |
| MeSH | D020121 |
| Anatomical terminology | |
The 5′ UTR has been found to interact with proteins relating to metabolism, and proteins translate sequences within the 5′ UTR. In addition, this region has been involved in transcription regulation, such as the sex-lethal gene in Drosophila.[1] Regulatory elements within 5′ UTRs have also been linked to mRNA export.[2]
General structure
Length
The 5′ UTR begins at the transcription start site and ends one nucleotide (nt) before the initiation sequence (usually AUG) of the coding region. In prokaryotes, the length of the 5′ UTR tends to be 3–10 nucleotides long, while in eukaryotes it tends to be anywhere from 100 to several thousand nucleotides long.[3] For example, the ste11 transcript in Schizosaccharomyces pombe has a 2273 nucleotide 5′ UTR[4] while the lac operon in Escherichia coli only has seven nucleotides in its 5′ UTR.[5] The differing sizes are likely due to the complexity of the eukaryotic regulation which the 5′ UTR holds as well as the larger pre-initiation complex that must form to begin translation.
The 5′ UTR can also be completely missing, in the case of leaderless mRNAs. Ribosomes of all three domains of life accept and translate such mRNAs.[6] Such sequences are naturally found in all three domains of life. Humans have many pressure-related genes under a 2–3 nucleotide leader. Mammals also have other types of ultra-short leaders like the TISU sequence.[7]
Elements
The elements of a eukaryotic and prokaryotic 5′ UTR differ greatly. The prokaryotic 5′ UTR contains a ribosome binding site (RBS), also known as the Shine–Dalgarno sequence (AGGAGGU), which is usually 3–10 base pairs upstream from the initiation codon.[5] In contrast, the eukaryotic 5′ UTR contains the Kozak consensus sequence (ACCAUGG), which contains the initiation codon.[5] The eukaryotic 5′ UTR also contains cis-acting regulatory elements called upstream open reading frames (uORFs) and upstream AUGs (uAUGs) and termination codons, which have a great impact on the regulation of translation (see below). Unlike prokaryotes, 5′ UTRs can harbor introns in eukaryotes. In humans, ~35% of all genes harbor introns within the 5′ UTR.[8]
Secondary structure
As the 5′ UTR has high GC content, secondary structures often occur within it. Hairpin loops are one such secondary structure that can be located within the 5′ UTR. These secondary structures also impact the regulation of translation.[9]
Role in translational regulation
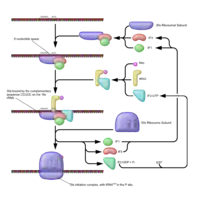
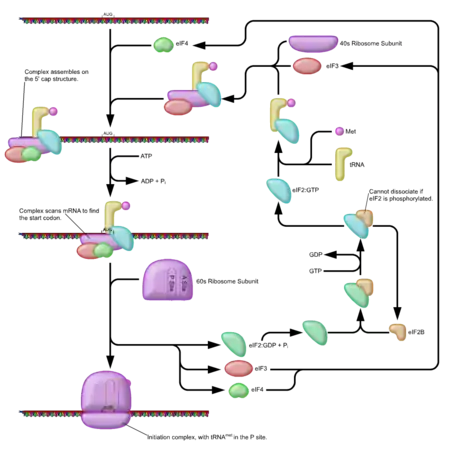
Prokaryotes
In bacteria, the initiation of translation occurs when IF-3, along with the 30S ribosomal subunit, bind to the Shine–Dalgarno (SD) sequence of the 5′ UTR.[5] This then recruits many other proteins, such as the 50S ribosomal subunit, which allows for translation to begin. Each of these steps regulates the initiation of translation.
Initiation in Archaea is less understood. SD sequences are much rarer, and the initiation factors have more in common with eukaryotic ones. There is no homolog of bacterial IF3.[10] Some mRNAs are leaderless.[11]
In both domains, genes without Shine–Dalgarno sequences are also translated in a less understood manner. A requirement seems to be a lack of secondary structure near the initiation codon.[12]
Pre-initiation complex regulation
The regulation of translation in eukaryotes is more complex than in prokaryotes. Initially, the eIF4F complex is recruited to the 5′ cap, which in turn recruits the ribosomal complex to the 5′ UTR. Both eIF4E and eIF4G bind the 5′ UTR, which limits the rate at which translational initiation can occur. However, this is not the only regulatory step of translation that involves the 5′ UTR.
RNA-binding proteins sometimes serve to prevent the pre-initiation complex from forming. An example is regulation of the msl2 gene. The protein SXL attaches to an intron segment located within the 5′ UTR segment of the primary transcript, which leads to the inclusion of the intron after processing.[13] This sequence allows the recruitment of proteins that bind simultaneously to both the 5′ and 3′ UTR, not allowing translation proteins to assemble. However, it has also been noted that SXL can also repress translation of RNAs that do not contain a poly(A) tail, or more generally, 3′ UTR.

Closed-loop regulation
Another important regulator of translation is the interaction between 3′ UTR and the 5′ UTR.
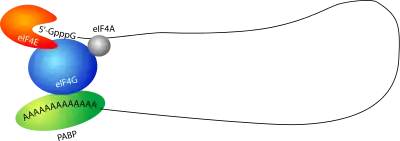
The closed-loop structure inhibits translation. This has been observed in Xenopus laevis, in which eIF4E bound to the 5′ cap interacts with Maskin bound to CPEB on the 3′ UTR, creating translationally inactive transcripts. This translational inhibition is lifted once CPEB is phosphorylated, displacing the Maskin binding site, allowing for the polymerization of the PolyA tail, which can recruit the translational machinery by means of PABP.[14] However, it is important to note that this mechanism has been under great scrutiny.[15]
Ferritin regulation
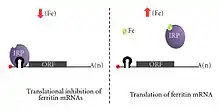
Iron levels in cells are maintained by translation regulation of many proteins involved in iron storage and metabolism. The 5′ UTR has the ability to form a hairpin loop secondary structure (known as the iron response element or IRE) that is recognized by iron-regulatory proteins (IRP1 and IRP2). In low levels of iron, the ORF of the target mRNA is blocked as a result of steric hindrance from the binding of IRP1 and IRP2 to the IRE. When iron is high, then the two iron-regulatory proteins do not bind as strongly and allow proteins to be expressed that have a role in iron concentration control. This function has gained some interest after it was revealed that the translation of amyloid precursor protein may be disrupted due to a single-nucleotide polymorphism to the IRE found in the 5′ UTR of its mRNA, leading to a spontaneous increased risk of Alzheimer's disease.[16]
uORFs and reinitiation
Another form of translational regulation in eukaryotes comes from unique elements on the 5′ UTR called upstream open reading frames (uORF). These elements are fairly common, occurring in 35–49% of all human genes.[17] A uORF is a coding sequence located in the 5′ UTR located upstream of the coding sequences initiation site. These uORFs contain their own initiation codon, known as an upstream AUG (uAUG). This codon can be scanned for by ribosomes and then translated to create a product,[18] which can regulate the translation of the main protein coding sequence or other uORFs that may exist on the same transcript.
The translation of the protein within the main ORF after a uORF sequence has been translated is known as reinitiation.[19] The process of reinitiation is known to reduce the translation of the ORF protein. Control of protein regulation is determined by the distance between the uORF and the first codon in the main ORF.[19] A uORF has been found to increase reinitiation with the longer distance between its uAUG and the start codon of the main ORF, which indicates that the ribosome needs to reacquire translation factors before it can carry out translation of the main protein.[19] For example, ATF4 regulation is performed by two uORFs further upstream, named uORF1 and uORF2, which contain three amino acids and fifty-nine amino acids, respectively. The location of uORF2 overlaps with the ATF4 ORF. During normal conditions, the uORF1 is translated, and then translation of uORF2 occurs only after eIF2-TC has been reacquired. Translation of the uORF2 requires that the ribosomes pass by the ATF4 ORF, whose start codon is located within uORF2. This leads to its repression. However, during stress conditions, the 40S ribosome will bypass uORF2 because of a decrease in concentration of eIF2-TC, which means the ribosome does not acquire one in time to translate uORF2. Instead, ATF4 is translated.[19]
Other mechanisms
In addition to reinitiation, uORFs contribute to translation initiation based on:
- The nucleotides of an uORF may code for a codon that leads to a highly structured mRNA, causing the ribosome to stall.[19]
- cis- and trans- regulation on translation of the main protein coding sequence.[19]
- Interactions with IRES sites.[19]

Internal ribosome entry sites and viruses
Viral (as well as some eukaryotic) 5′ UTRs contain internal ribosome entry sites, which is a cap-independent method of translational activation. Instead of building up a complex at the 5′ cap, the IRES allows for direct binding of the ribosomal complexes to the transcript to begin translation.[20] The IRES enables the viral transcript to translate more efficiently due to the lack of needing a preinitation complex, allowing the virus to replicate quickly.[5]
Role in transcriptional regulation
msl-2 transcript
Transcription of the msl-2 transcript is regulated by multiple binding sites for fly Sxl at the 5′ UTR.[1] In particular, these poly-uracil sites are located close to a small intron that is spliced in males, but kept in females through splicing inhibition. This splicing inhibition is maintained by Sxl.[1] When present, Sxl will repress the translation of msl2 by increasing translation of a start codon located in a uORF in the 5′ UTR (see above for more information on uORFs). Also, Sxl outcompetes TIA-1 to a poly(U) region and prevents snRNP (a step in alternative splicing) recruitment to the 5′ splice site.[1]
See also
References
- Penalva, L. O. F.; Sanchez, L. (2003). "RNA Binding Protein Sex-Lethal (Sxl) and Control of Drosophila Sex Determination and Dosage Compensation". Microbiology and Molecular Biology Reviews. 67 (3): 343–59, table of contents. doi:10.1128/MMBR.67.3.343-359.2003. PMC 193869. PMID 12966139.
- Cenik, Can; Chua, Hon Nian; Zhang, Hui; Tarnawsky, Stefan P.; Akef, Abdalla; Derti, Adnan; Tasan, Murat; Moore, Melissa J.; Palazzo, Alexander F.; Roth, Frederick P. (2011). Snyder, Michael (ed.). "Genome Analysis Reveals Interplay between 5′UTR Introns and Nuclear mRNA Export for Secretory and Mitochondrial Genes". PLOS Genetics. 7 (4): e1001366. doi:10.1371/journal.pgen.1001366. ISSN 1553-7404. PMC 3077370. PMID 21533221.
- Lodish, Havery (2004). Molecular Cell Biology. New York, New York: W.H. Freeman and Company. p. 113. ISBN 978-0-7167-4366-8.
- Rhind, Nicholas; Chen, Zehua; Yassour, Moran; Thompson, Dawn A.; Haas, Brian J.; Habib, Naomi; Wapinski, Ilan; Roy, Sushmita; Lin, Michael F.; Heiman, David I.; Young, Sarah K.; Furuya, Kanji; Guo, Yabin; Pidoux, Alison; Chen, Huei Mei; Robbertse, Barbara; Goldberg, Jonathan M.; Aoki, Keita; Bayne, Elizabeth H.; Berlin, Aaron M.; Desjardins, Christopher A.; Dobbs, Edward; Dukaj, Livio; Fan, Lin; Fitzgerald, Michael G.; French, Courtney; Gujja, Sharvari; Hansen, Klavs; Keifenheim, Dan; Levin, Joshua Z. (2011). "Comparative Functional Genomics of the Fission Yeasts". Science. 332 (6032): 930–6. Bibcode:2011Sci...332..930R. doi:10.1126/science.1203357. PMC 3131103. PMID 21511999.
- Brown, T.A (2007). Genomes 3. New York, New York: Garland Science Publishing. p. 397. ISBN 978-0-8153-4138-3.
- Brock, JE; Pourshahian, S; Giliberti, J; Limbach, PA; Janssen, GR (October 2008). "Ribosomes bind leaderless mRNA in Escherichia coli through recognition of their 5'-terminal AUG". RNA. 14 (10): 2159–69. doi:10.1261/rna.1089208. PMC 2553737. PMID 18755843.
- Akulich, Kseniya A.; Andreev, Dmitry E.; Terenin, Ilya M.; Smirnova, Victoria V.; Anisimova, Aleksandra S.; Makeeva, Desislava S.; Arkhipova, Valentina I.; Stolboushkina, Elena A.; Garber, Maria B.; Prokofjeva, Maria M.; Spirin, Pavel V.; Prassolov, Vladimir S.; Shatsky, Ivan N.; Dmitriev, Sergey E. (28 November 2016). "Four translation initiation pathways employed by the leaderless mRNA in eukaryotes". Scientific Reports. 6 (1): 37905. Bibcode:2016NatSR...637905A. doi:10.1038/srep37905. PMC 5124965. PMID 27892500.
- Bicknell AA, Cenik C, Chua HN, Roth FP, Moore MJ (Dec 2012). "Introns in UTRs: why we should stop ignoring them". BioEssays. 34 (12): 1025–34. doi:10.1002/bies.201200073. PMID 23108796.
- Babendure, J. R.; Babendure, JL; Ding, JH; Tsien, RY (2006). "Control of mammalian translation by mRNA structure near caps". RNA. 12 (5): 851–61. doi:10.1261/rna.2309906. PMC 1440912. PMID 16540693.
- Benelli, D; Londei, P (January 2011). "Translation initiation in Archaea: conserved and domain-specific features". Biochemical Society Transactions. 39 (1): 89–93. doi:10.1042/BST0390089. PMID 21265752.
- Hernández, Greco; Jagus, Rosemary (2016-08-10). "Evolution of Translational Initiation: From Archaea to Eukarya". Evolution of the protein synthesis machinery and its regulation. Hernández, Greco,, Jagus, Rosemary. Switzerland. doi:10.1007/978-3-319-39468-8_4. ISBN 9783319394688. OCLC 956539514.
- Nakagawa, S; Niimura, Y; Gojobori, T (20 April 2017). "Comparative genomic analysis of translation initiation mechanisms for genes lacking the Shine-Dalgarno sequence in prokaryotes". Nucleic Acids Research. 45 (7): 3922–3931. doi:10.1093/nar/gkx124. PMC 5397173. PMID 28334743.
- Araujo, Patricia R.; Yoon, Kihoon; Ko, Daijin; Smith, Andrew D.; Qiao, Mei; Suresh, Uthra; Burns, Suzanne C.; Penalva, Luiz O. F. (2012). "Before It Gets Started: Regulating Translation at the 5′ UTR". Comparative and Functional Genomics. 2012: 1–8. doi:10.1155/2012/475731. PMC 3368165. PMID 22693426.
- Gilbert, Scott (2010). Developmental Biology. Sunderland, MA: Sinauer Associates. p. 60. ISBN 978-0-87893-384-6.
- Kozak, Marilyn (2008). "Faulty old ideas about translational regulation paved the way for current confusion about how microRNAs function". Gene. 423 (2): 108–15. doi:10.1016/j.gene.2008.07.013. PMID 18692553.
- Rogers, Jack T.; Bush, Ashley I.; Cho, Hyan-Hee; Smith, Deborah H.; Thomson, Andrew M.; Friedlich, Avi L.; Lahiri, Debomoy K.; Leedman, Peter J.; Huang, Xudong; Cahill, Catherine M. (2008). "Iron and the translation of the amyloid precursor protein (APP) and ferritin mRNAs: Riboregulation against neural oxidative damage in Alzheimer's disease". Biochemical Society Transactions. 36 (6): 1282–7. doi:10.1042/BST0361282. PMC 2746665. PMID 19021541.
- Mignone, Flavio; Gissi, Carmela; Liuni, Sabino; Pesole, Graziano (2002). "Untranslated regions of mRNAs". Genome Biology. 3 (3): reviews0004.1. doi:10.1186/gb-2002-3-3-reviews0004. PMC 139023. PMID 11897027.
- Wethmar, Klaus; Smink, Jeske J.; Leutz, Achim (2010). "Upstream open reading frames: Molecular switches in (patho)physiology". BioEssays. 32 (10): 885–93. doi:10.1002/bies.201000037. PMC 3045505. PMID 20726009.
- Somers, Joanna; Pöyry, Tuija; Willis, Anne E. (2013). "A perspective on mammalian upstream open reading frame function". The International Journal of Biochemistry & Cell Biology. 45 (8): 1690–700. doi:10.1016/j.biocel.2013.04.020. PMC 7172355. PMID 23624144.
- Thompson, Sunnie R. (2012). "Tricks an IRES uses to enslave ribosomes". Trends in Microbiology. 20 (11): 558–66. doi:10.1016/j.tim.2012.08.002. PMC 3479354. PMID 22944245.