Foldit
Foldit is an online puzzle video game about protein folding. It is part of an experimental research project developed by the University of Washington, Center for Game Science, in collaboration with the UW Department of Biochemistry. The objective of Foldit is to fold the structures of selected proteins as perfectly as possible, using tools provided in the game. The highest scoring solutions are analyzed by researchers, who determine whether or not there is a native structural configuration (native state) that can be applied to relevant proteins in the real world. Scientists can then use these solutions to target and eradicate diseases and create biological innovations. A 2010 paper in the science journal Nature credited Foldit's 57,000 players with providing useful results that matched or outperformed algorithmically computed solutions.[3][4]
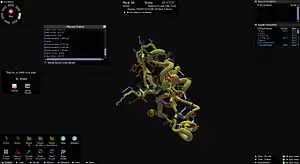 Screenshot of the Foldit application showing a folding puzzle in progress | |
| Developer(s) | University of Washington, Center for Game Science,[1] Department of Biochemistry[2] |
|---|---|
| Initial release | May 8, 2008 |
| Preview release | |
| Operating system | Cross-platform: Windows, macOS, Linux |
| Size | ≈434 MB |
| Available in | 13 languages |
List of languages Czech, Dutch, English, French, German, Hebrew, Indonesian, Italian, Polish, Romanian, Russian, Spanish, Scientist | |
| Type | Puzzle video game, protein folding |
| License | proprietary freeware for academic and non-profit use |
| Website | fold |
History
Rosetta
Prof. David Baker, a protein research scientist at the University of Washington, founded the Foldit project. Seth Cooper was the lead game designer. Before starting the project, Baker and his laboratory coworkers relied on another research project named Rosetta[5] to predict the native structures of various proteins using special computer protein structure prediction algorithms. Rosetta was eventually extended to use the power of distributed computing: The Rosetta@home program was made available for public download, and displayed its protein-folding progress as a screensaver. Its results were sent to a central server for verification.[6]
Some Rosetta@home users became frustrated when they saw ways to solve protein structures, but could not interact with the program. Hoping that humans could improve the computers' attempts to solve protein structures, Baker approached David Salesin and Zoran Popović, computer science professors at the same university, to help conceptualize and build an interactive program, a video game, that would appeal to the public and help efforts to find native protein structures.[7][8][9]
Foldit
Many of the same people who created Rosetta@home worked on Foldit. The public beta version was released in May 2008[10] and has 240,000 registered players.[11]
Since 2008, Foldit has participated in Critical Assessment of Techniques for Protein Structure Prediction (CASP) experiments, submitting its best solutions to targets based on unknown protein structures. CASP is an international program to assess methods of protein structure prediction and identify those that are most productive.
Goals
Protein structure prediction is important in several fields of science, including bioinformatics, molecular biology, and medicine. Identifying natural proteins' structural configurations enables scientists to understand them better. This can lead to creating novel proteins by design, advances in treating disease, and solutions for other real-world problems such as invasive species, waste, and pollution.
The process by which living beings create the primary structure of proteins, protein biosynthesis, is reasonably well understood, as is the means by which proteins are encoded as DNA. However, determining how a given protein's primary structure becomes a functioning three-dimensional structure, how the molecule folds, is more difficult. The general process is understood, but predicting a protein's eventual, functioning structure is computationally demanding.[12][13]
Methods
Similarly to Rosetta@home, Foldit is a means to discover native protein structures faster through distributed computing. However, Foldit has a greater emphasis on community collaboration through its forums, where users can collaborate on certain folds.[3] Furthermore, Foldit's crowdsourced approach places a greater emphasis on the user.[6] Foldit's virtual interaction and gamification create a unique and innovative environment with the potential to greatly advance protein folding research.
Virtual interaction
Foldit attempts to apply the human brain's three-dimensional pattern matching and spatial reasoning abilities to help solve the problem of protein structure prediction. 2016 puzzles are based on well-understood proteins. By analysing how humans intuitively approach these puzzles, researchers hope to improve the algorithms used by protein-folding software.[14]
Foldit includes a series of tutorials where users manipulate simple protein-like structures and a periodically updated set of puzzles based on real proteins. It shows a graphical representation of each protein which users can manipulate using a set of tools.
Gamification
Foldit's developers wanted to attract as many people as possible to the cause of protein folding. So, rather than only building a useful science tool, they used gamification (the inclusion of gaming elements) to make Foldit appealing and engaging to the general public.
As a protein structure is modified, a score is calculated based on how well-folded the protein is, and a list of high scores for each puzzle is maintained. Foldit users may create and join groups, and members of groups can share puzzle solutions. Groups have been found to be useful in training new players. A separate list of group high scores is maintained.
Accomplishments
Results from Foldit have been included in a number of scientific publications.
Foldit players have been cited collectively as "Foldit players" or "Players, F." in some cases. Individual players have also been listed as authors on at least one paper, and on four related Protein Data Bank depositions.
- An August 2010 paper in the journal Nature credited Foldit's 57,000 players with providing useful results that matched or outperformed algorithmically computed solutions, stating "[p]layers working collaboratively develop a rich assortment of new strategies and algorithms; unlike computational approaches, they explore not only conformational space but also the space of possible search strategies".[15][3]
- A November 2011 article in PNAS compared "recipes" developed by Foldit players to Rosetta scripts developed by members of the Baker Lab at the University of Washington. The player-developed "Blue Fuse" recipe compared favorably with the scientists' "Fast Relax" algorithm.[16]
- In 2011, Foldit players helped decipher the crystal structure of a retroviral protease from Mason-Pfizer monkey virus (M-PMV), a monkey virus which causes HIV/AIDS-like symptoms, a scientific problem that had been unsolved for 15 years. While the puzzle was available for three weeks, players produced a 3D model of the enzyme in only ten days that is accurate enough for molecular replacement.[17][18][19]
- In January 2012, Scientific American reported that Foldit gamers achieved the first crowdsourced redesign of a protein,[11] an enzyme that catalysed the Diels–Alder reactions widely used in synthetic chemistry. A team including David Baker in the Center for Game Science at University of Washington in Seattle computationally designed the enzyme from scratch but found its potency needed improvement. Foldit players reengineered the enzyme by adding 13 amino acids, increasing its activity by more than 18 times.[11][20]
- A September 2016 article in Nature Communications detailed a "crystallographic model-building competition between trained crystallographers, undergraduate students, Foldit players and automatic model-building algorithms" in which "a team of Foldit players achieved the most accurate structure" fitting a protein to the results of an X-ray crystallography experiment.[21]
- A July 2018 article in Nature Communications reviewed the collaboration between Foldit players and teams in the WeFold consortium in biennial CASP competitions CASP11 and CASP12.[22]
- A June 2019 letter in Nature described the analysis of proteins designed by Foldit players. Four player-designed proteins were successfully grown in E. coli and then "solved" via X-ray crystallography. The proteins were added to the Protein Data Bank as 6MRR, 6MRS, 6MSP, and 6NUK.[23]
- In November 2019, an article in PLOS Biology reported how Foldit players were able to "build protein structures into crystallographic, high-resolution maps more accurately than expert crystallographers or automated model-building algorithms" using data from cryo EM experiments.[24]
Future development
Foldit's toolbox is mainly for the design of protein molecules. The game's creator announced the plan to add, by 2013, the chemical building blocks of organic subcomponents to enable players to design small molecules.[25]
See also
References
- University of Washington, Center for Game Science
- University of Washington, Department of Biochemistry
- Markoff J (10 August 2010). "In a Video Game, Tackling the Complexities of Protein Folding". The New York Times. Retrieved 12 February 2013.
- Cooper S, Khatib F, Treuille A, Barbero J, Lee J, Beenen M, et al. (August 2010). "Predicting protein structures with a multiplayer online game". Nature. 466 (7307): 756–60. Bibcode:2010Natur.466..756C. doi:10.1038/nature09304. PMC 2956414. PMID 20686574.
- "Rosetta Commons: The hub for Rosetta modeling software". RosettaCommons.org. RosettaCommons.org. Retrieved 17 November 2015.
- Howard Hughes Medical Institute "Protein-folding game taps power of worldwide audience to solve difficult puzzles" Eurekalert!, August 4, 2010
- Bourzac K (2008-05-08). "Biologists Enlist Online Gamers". Technology Review. Retrieved 17 November 2016.
- Bohannon J (2009-04-20). "Gamers Unravel the Secret Life of Protein". Wired. Retrieved 17 November 2016.
- "Zoran Popović". washington.edu.
- Hickey, Hannah. "Computer game's high score could earn the Nobel Prize in medicine" University of Washington, May 8, 2008
- Marshall J (January 22, 2012). "Online Gamers Achieve First Crowd-Sourced Redesign of Protein". Scientific American. Retrieved February 22, 2012.
- Haspel N, Tsai CJ, Wolfson H, Nussinov R (June 2003). "Reducing the computational complexity of protein folding via fragment folding and assembly". Protein Science. 12 (6): 1177–87. doi:10.1110/ps.0232903. PMC 2323902. PMID 12761388.
- Rocklin GJ, Chidyausiku TM, Goreshnik I, Ford A, Houliston S, Lemak A, et al. (July 2017). "Global analysis of protein folding using massively parallel design, synthesis, and testing". Science. 357 (6347): 168–175. Bibcode:2017Sci...357..168R. doi:10.1126/science.aan0693. PMC 5568797. PMID 28706065.
- Horowitz S, Koepnick B, Martin R, Tymieniecki A, Winburn AA, Cooper S, et al. (September 2016). "Determining crystal structures through crowdsourcing and coursework". Nature Communications. 7: 12549. Bibcode:2016NatCo...712549H. doi:10.1038/ncomms12549. PMC 5028414. PMID 27633552.
- Cooper S, Khatib F, Treuille A, Barbero J, Lee J, Beenen M, et al. (August 2010). "Predicting protein structures with a multiplayer online game". Nature. 466 (7307): 756–60. Bibcode:2010Natur.466..756C. doi:10.1038/nature09304. PMC 2956414. PMID 20686574.
- Khatib F, Cooper S, Tyka MD, Xu K, Makedon I, Popovic Z, et al. (November 2011). "Algorithm discovery by protein folding game players". Proceedings of the National Academy of Sciences of the United States of America. 108 (47): 18949–53. doi:10.1073/pnas.1115898108. PMC 3223433. PMID 22065763.
- Khatib F, DiMaio F, Cooper S, Kazmierczyk M, Gilski M, Krzywda S, et al. (September 2011). "Crystal structure of a monomeric retroviral protease solved by protein folding game players". Nature Structural & Molecular Biology. 18 (10): 1175–7. doi:10.1038/nsmb.2119. PMC 3705907. PMID 21926992.
- Gilski M, Kazmierczyk M, Krzywda S, Zábranská H, Cooper S, Popović Z, et al. (November 2011). "High-resolution structure of a retroviral protease folded as a monomer". Acta Crystallographica. Section D, Biological Crystallography. 67 (Pt 11): 907–14. doi:10.1107/S0907444911035943. PMC 3211970. PMID 22101816.
- Praetorius D (2011-09-19). "Gamers Decode AIDS Protein That Stumped Researchers For 15 Years In Just 3 Weeks". The Huffington Post. Retrieved 17 November 2016.
- Eiben CB, Siegel JB, Bale JB, Cooper S, Khatib F, Shen BW, et al. (January 2012). "Increased Diels-Alderase activity through backbone remodeling guided by Foldit players". Nature Biotechnology. 30 (2): 190–2. doi:10.1038/nbt.2109. PMC 3566767. PMID 22267011.
- Horowitz S, Koepnick B, Martin R, Tymieniecki A, Winburn AA, Cooper S, et al. (September 2016). "Determining crystal structures through crowdsourcing and coursework". Nature Communications. 7 (1): 12549. Bibcode:2016NatCo...712549H. doi:10.1038/ncomms12549. PMID 27633552.
- Keasar C, McGuffin LJ, Wallner B, Chopra G, Adhikari B, Bhattacharya D, et al. (July 2018). "An analysis and evaluation of the WeFold collaborative for protein structure prediction and its pipelines in CASP11 and CASP12". Scientific Reports. 8 (1): 9939. Bibcode:2018NatSR...8.9939K. doi:10.1038/s41598-018-26812-8. PMC 6028396. PMID 29967418.
- Koepnick B, Flatten J, Husain T, Ford A, Silva DA, Bick MJ, et al. (June 2019). "De novo protein design by citizen scientists". Nature. 570 (7761): 390–394. Bibcode:2019Natur.570..390K. doi:10.1038/s41586-019-1274-4. PMC 6701466. PMID 31168091.
- Khatib F, Desfosses A, Koepnick B, Flatten J, Popović Z, Baker D, et al. (November 2019). "Building de novo cryo-electron microscopy structures collaboratively with citizen scientists". PLOS Biology. 17 (11): e3000472. doi:10.1371/journal.pbio.3000472. PMC 6850521. PMID 31714936.
- Hersher R (April 13, 2012). "FoldIt game's next play: crowdsourcing better drug design". nature.com. Retrieved April 16, 2012.
External links
- official Foldit website
- Foldit wiki
- Baker Lab at the University of Washington
- Institute for Protein Design
- Center for Game Science