Gene map
Gene maps help describe the spatial arrangement of genes on a chromosome. Genes are designated to a specific location on a chromosome known as the locus and can be used as molecular markers to find the distance between other genes on a chromosome. Maps provide researchers with the opportunity to predict the inheritance patterns of specific traits, which can eventually lead to a better understanding of disease-linked traits.[1]
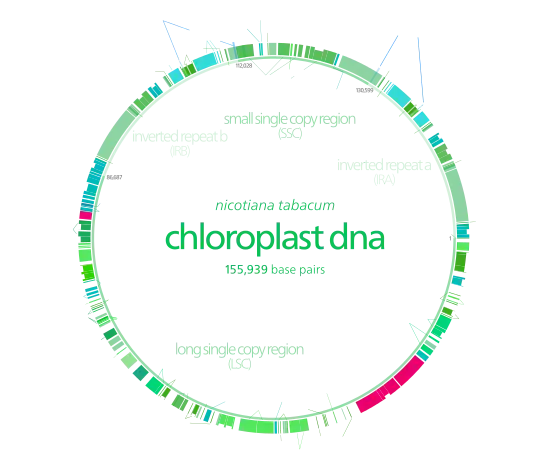
The genetic basis to gene maps is to provide an outline that can potentially help researchers carry out DNA sequencing. A gene map helps point out the relative positions of genes and allows researchers to locate regions of interest in the genome. Genes can then be identified quickly and sequenced quickly.[2]
Two approaches to generating gene maps include physical mapping and genetic mapping. Physical mapping utilizes molecular biology techniques to inspect chromosomes. These techniques consequently allow researchers to observe chromosomes directly so that a map may be constructed with relative gene positions. Genetic mapping on the other hand uses genetic techniques to indirectly find association between genes. Techniques can include cross-breeding (see Hybrid (biology)) experiments and examining pedigrees. These technique allow for maps to be constructed so that relative positions of genes and other important sequences can be analyzed.[2]
Physical mapping
Physical mapping techniques used to generate a gene map include: Restriction mapping, Fluorescent in situ hybridization (FISH), and Sequence tagged site (STS) mapping.
Restriction mapping
Restriction mapping is a method in which structural information regarding a segment of DNA is obtained using restriction enzymes. Restriction enzymes are enzymes that help cut segments of DNA at specific recognition sequences. The basis to restriction mapping involves digesting (or cutting) DNA with restriction enzymes. The digested DNA fragments are then run on an agarose gel using electrophoresis, which provides one with information regarding the size of these digested fragments. The sizes of these fragments help indicate the distance between restriction enzyme sites on the DNA analyzed, and provides researchers with information regarding the structure of DNA analyzed.[3]
Fluorescent in situ hybridization (FISH)
FISH is a method used to detect the presence (or absence) of a DNA sequence within a cell. DNA probes that are specific for chromosomal regions or genes of interest are labeled with fluorochromes. By attaching fluorochromes to probes, researchers are able to visualize multiple DNA sequences simultaneously. When a probe comes into contact with DNA on a specific chromosome, hybridization will occur. Consequently, information regarding the location of that sequence of DNA will be attained. FISH analyzes single stranded DNA (ssDNA). Once the DNA is in its single stranded state, the DNA can bind to its specific probe.[2]
Sequence-tagged site (STS) mapping
A sequence-tagged site (STS) is a short sequence of DNA (about 100 - 500 base pairs in length) that is seen to appear multiple times within an individual's genome. These sites are easily recognizable, usually appearing at least once in the DNA being analyzed. These sites usually contain genetic polymorphisms making them sources of viable genetic markers (as they differ from other sequences). Sequenced tagged sites can be mapped within our genome and require a group of overlapping DNA fragments. PCR is generally used to produce the collection of DNA fragments. After overlapping fragments are created, the map distance between STSs can be analyzed. In order to calculate the map distance between STSs, researchers determine the frequency at which breaks between the two markers occur (see shotgun sequencing)[3]
Genetic mapping
Genetic mapping is focused on the principles first established by Gregor Mendel. This approach primarily focuses on linkage analysis and gene association techniques.
Linkage analysis
The basis to linkage analysis is understanding chromosomal location and identifying disease genes. Certain genes that are linked or associated with each other are found to reside close to each other on the same chromosome. During meiosis, these genes are capable of being inherited together and can be used as a genetic marker to help identify the phenotype of diseases. Because linkage analysis can identify inheritance patterns, these studies are usually family based.[4]
Gene association analysis
Gene association analysis is population based; it is not focused on inheritance patterns, but rather is based on the entire history of a population. Gene association analysis looks at a particular population and tries to identify whether the frequency of an allele in affected individuals is different from that of a control set of unaffected individuals of the same population. This method is particularly useful to identify complex diseases that do not have a Mendelian inheritance pattern.[3]
Utilizing gene maps: disease genes
Using the methods mentioned above, researchers are capable of mapping disease genes. Generating a gene map is the critical first step towards identifying disease genes. Gene maps allow for variant alleles to be identified and allow for researchers to make predictions about the genes they think are causing the mutant phenotype. An example of a disorder that was identified by Linkage analysis is Cystic Fibrosis. For example, with Cystic Fibrosis (CF), DNA samples from fifty families affected by CF were analyzed using linkage analysis. Hundreds of markers pertaining to CF were analyzed throughout the genome until CF was identified on the long arm of chromosome 7. Researchers then had completed linkage analysis on additional DNA markers within chromosome 7 to identify an even more precise location of the CF gene. They found that the CF gene resides around 7q31-q32 (see chromosomal nomenclature).[3]
See also
References
- Nussbaum, Robert L.; McInnes, Roderick R.; Wilard, Huntington F. (2016). Thompson & Thompson Genetics in Medicine (Eighth ed.). Philadelphia, PA: Elsevier. pp. 178–187. ISBN 978-1-4377-0696-3. Archived from the original on 4 March 2016. Retrieved 13 October 2015.
- Brown, Terence, A. (2002). Genomes. Manchester, UK: Garland Science. ISBN 0-471-25046-5.
- Hartwell, Leland H.; Hood, Leroy; Goldberg, Michael L.; Reynolds, Anne E.; Silver, Lee M.; Karagiannis, Jim; Papaconstantinou, Maria (2014). Genetics: From Genes to Genomes (Canadian ed.). Canada: McGraw-Hill Ryerson. pp. 456–459, 635–636. ISBN 978-0-07-094669-9. Retrieved 13 October 2015.
- Pulst, Stefan M. (June 1999). "Genetic Linkage Analysis". JAMA Neurology. 56 (6): 667–672. doi:10.1001/archneur.56.6.667. PMID 10369304. Retrieved 13 October 2015.
