Lactococcus lactis
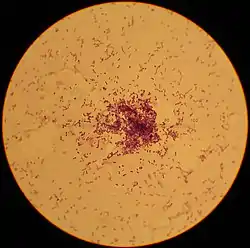
Lactococcus lactis is a Gram-positive bacterium used extensively in the production of buttermilk and cheese,[1] but has also become famous as the first genetically modified organism to be used alive for the treatment of human disease.[2] L. lactis cells are cocci that group in pairs and short chains, and, depending on growth conditions, appear ovoid with a typical length of 0.5 - 1.5 µm. L. lactis does not produce spores (nonsporulating) and are not motile (nonmotile). They have a homofermentative metabolism, meaning they produce lactic acid from sugars. They've also been reported to produce exclusive L-(+)-lactic acid.[3] However,[4] reported D-(−)-lactic acid can be produced when cultured at low pH. The capability to produce lactic acid is one of the reasons why L. lactis is one of the most important microorganisms in the dairy industry.[5] Based on its history in food fermentation, L. lactis has generally recognized as safe (GRAS) status,[6][7] with few case reports of it being an opportunistic pathogen.[8][9][10]
| "Lactobacillus lactis" | |
|---|---|
 | |
| Scientific classification | |
| Domain: | |
| Kingdom: | |
| Phylum: | |
| Class: | |
| Order: | |
| Family: | |
| Genus: | |
| Species: | L. lactis |
| Binomial name | |
| Lactococcus lactis (Lister 1873) Schleifer et al. 1986 | |
| Subspecies | |
|
L. l. cremoris | |
L. lactis is of crucial importance for manufacturing dairy products, such as buttermilk and cheeses. When L. lactis ssp. lactis is added to milk, the bacterium uses enzymes to produce energy molecules (ATP), from lactose. The byproduct of ATP energy production is lactic acid. The lactic acid produced by the bacterium curdles the milk, which then separates to form curds that are used to produce cheese.[11] Other uses that have been reported for this bacterium include the production of pickled vegetables, beer or wine, some breads, and other fermented foodstuffs like soymilk kefir, buttermilk, and others.[12] L. lactis is one of the best characterized low GC Gram positive bacteria with detailed knowledge on genetics, metabolism and biodiversity.[13][14]
L. lactis is mainly isolated from either the dairy environment, or plant material.[15][16][17] Dairy isolates are suggested to have evolved from plant isolates through a process in which genes without benefit in the rich milk were lost, or down-regulated.[14][18] This process, also called genome erosion or reductive evolution is also described in several other lactic acid bacteria.[19][20] The proposed transition from the plant to the dairy environment was reproduced in the laboratory through experimental evolution of a plant isolate that was cultivated in milk for a prolonged period. Consistent with the results from comparative genomics (see references above) this resulted in L. lactis losing or down-regulating genes which are dispensable in milk and the up-regulation of peptide transport.[21]
Hundreds of novel small RNAs were identified by Meulen et al. in the genome of L. lactis MG1363. One of them: LLnc147, was shown to be involved in carbon uptake and metabolism.[22]
Cheese production
L. lactis subsp. lactis (formerly Streptococcus lactis)[23] is used in the early stages for the production of many cheeses, including brie, camembert, Cheddar, Colby, Gruyère, Parmesan, and Roquefort.[24] The state Assembly of Wisconsin, also the number one cheese-producing state in the United States, voted in 2010 to name this bacterium as the official state microbe; it would have been the first and only such designation by a state legislature in the nation,[25] however the legislation was not adopted by the Senate.[26] The legislation was introduced in November 2009 as Assembly Bill 556 by Representatives Hebl, Vruwink, Williams, Pasch, Danou, and Fields; it was cosponsored by Senator Taylor.[27] The bill passed the Assembly on May 15, 2010, and was dropped by the Senate on April 28th.[27]
The use of L. lactis in dairy factories is not without issues. Bacteriophages specific to L. lactis cause significant economic losses each year by preventing the bacteria from fully metabolizing the milk substrate.[24] Several epidemiologic studies showed the phages mainly responsible for these losses are from the species 936, c2, and P335 (all from the family Siphoviridae).[28]
Therapeutic benefits
The feasibility of using Lactic acid bacteria (LAB) as functional protein delivery vectors has been widely investigated.[29] Lactococcus lactis has been demonstrated to be a promising candidate for the delivery of functional proteins because of its noninvasive and nonpathogenic characteristics.[30] Many different expression systems of L. lactis have been developed and used for heterologous protein expression.[31][32][33]
Lactose fermentation In Shuichi Nakamura’s, Yusuke V. Marimoto, and Seishi Kudo’s study, they sought to prove that some fermentation produced by L. lactis can hinder motility in pathogenic bacteria. The motilities of Pseudomonas, Vibrio, and Leptospira strains were also severely disrupted by lactose utilization by L. lactis.[34]
Using Salmonella flagellar as the experimental group, Nakamura’s team found that a product of lactose fermentation is the cause of motility impairment in Salmonella. It is suggested that the L. lactis supernatant mainly affects Salmonella motility through disturbing flagellar rotation but not through irreversible damage against morphologies and physiologies. Lactose fermentation by L. lactis produces acetate that reduces the intracellular pH of Salmonella, which in turn slows the rotation of their flagella.[35][36] These results highlight the potential use of L. lactis for preventing infections by multiple bacterial species.
Secretion of Interleukin-10 Genetically engineered L. lactis can secrete the cytokine interleukin-10 (IL-10) for the treatment of inflammatory bowel diseases (IBD), since IL-10 has a central role in downregulating inflammatory cascades[37] and matrix metalloproteinases.[38] A study by Lothar Steidler and Wolfgang Hans [39] shows that this in situ synthesis of IL-10 by genetically engineered L. lactis requires much lower doses than systemic treatments like antibodies to tumor necrosis factor (TNF) or recombination IL-10.
The authors propose two possible routes by which IL-10 can reach its therapeutic target. Genetically engineered L. lactis may produce murine IL-10 in the lumen, and the protein may diffuse to responsive cells in the epithelium or the lamina propria. Another route involves L. lactis taken up by M cells because of its bacterial size and shape, and the major part of the effect may be due to recombinant IL-10 production in situ in intestinal lymphoid tissue. Both routes may involve paracellular transport mechanisms that are enhanced in inflammation. After transport, IL-10 may directly down-regulate inflammation. In principle, this method may be useful for intestinal delivery of other protein therapeutics that are unstable or difficult to produce in large quantities and an alternative to the systemic treatment of IBD.
Tumor-suppressor through Tumor metastasis-inhibiting peptide KISS1 Another study, led by Zhang B, created a L. lactis strain that maintains a plasmid containing a tumor metastasis-inhibiting peptide known as KISS1.[40] L. lactis NZ9000 was demonstrated to be a cell factory for the secretion of biologically active KiSS1 protein, exerting inhibition effects on human colorectal cancer HT-29 cells.
KiSS1 secreted from recombinant L. lactis strain effectively downregulated the expression of Matrix metalloproteinases (MMP-9) – a crucial key in the invasion, metastasis, and regulation of the signaling pathways controlling tumor cell growth, survival, invasion, inflammation, and angiogenesis.[41][42][43] The reason for this is that KiSS1 expressed in L. lactis activates the MAPK pathway via GPR54 signaling, suppressing NFκB binding to the MMP-9 promoter and thus downregulating MMP-9 expression.[44] This, in turn, reduces the survival rate, inhibits metastasis and induces dormancy of cancer cells.
In addition, it was demonstrated that tumor growth can be inhibited by the LAB strain itself [45][46] due to the LAB’s ability to produce exopolysaccharides.[47][48] This study shows that L. lactisNZ9000 can inhibit HT-29 proliferation and induce cell apoptosis by itself. The success of this strain’s construction helped to inhibit migration and expansion of cancer cells, showing that the secretion properties of L. lactis of this particular peptide may serve as a new tool for cancer therapy in the future.[49]
References
- Madigan M, Martinko J (editors). (2005). Brock Biology of Microorganisms (11th ed.). Prentice Hall. ISBN 978-0-13-144329-7.
- Braat H, Rottiers P, Hommes DW, Huyghebaert N, Remaut E, Remon JP, van Deventer SJ, Neirynck S, Peppelenbosch MP, Steidler L (2006). "A phase I trial with transgenic bacteria expressing interleukin-10 in Crohn's disease". Clin Gastroenterol Hepatol. 4 (6): 754–759. doi:10.1016/j.cgh.2006.03.028. PMID 16716759.
- ROISSART, H. and Luquet F.M. Bactéries lactiques: aspects fondamentaux et technologiques. Uriage, Lorica, France, 1994, vol. 1, p. 605. ISBN 2-9507477-0-1
- Åkerberg, C.; Hofvendahl, K.; Zacchi, G.; Hahn-Hä;gerdal, B. (1998). "Modelling the influence of pH, temperature, glucose and lactic acid concentrations on the kinetics of lactic acid production by Lactococcus lactis ssp. Lactis ATCC 19435 in whole-wheat flour". Applied Microbiology and Biotechnology. 49 (6): 682–690. doi:10.1007/s002530051232. S2CID 46383610.CS1 maint: multiple names: authors list (link)
- Integr8 - Species search results:
- FDA. "History of the GRAS List and SCOGS Reviews". FDA. Retrieved 11 May 2012.
- Wessels S, Axelsson L, Bech Hansen E, De Vuyst L, Laulund S, Lähteenmäki L, Lindgren S, et al. (November 2004). "The lactic acid bacteria, the food chain, and their regulation". Trends in Food Science & Technology. 15 (10): 498–505. doi:10.1016/j.tifs.2004.03.003.
- Aguirre M, Collins MD (August 1993). "Lactic acid bacteria and human clinical infection". J. Appl. Bacteriol. 75 (2): 95–107. doi:10.1111/j.1365-2672.1993.tb02753.x. PMID 8407678.
- Facklam RR, Pigott NE, Collins MD. Identification of Lactococcus species from human sources. Proceedings of the XI Lancefield International Symposium on Streptococci and Streptococcal Diseases, Siena, Italy. Stuttgart: Gustav Fischer Verlag; 1990:127
- Mannion PT, Rothburn MM (November 1990). "Diagnosis of bacterial endocarditis caused by Streptococcus lactis and assisted by immunoblotting of serum antibodies". J. Infect. 21 (3): 317–8. doi:10.1016/0163-4453(90)94149-T. PMID 2125626.
- Lactococcus_lactis
- Lactococcus lactis uses
- Kok J, Buist G, Zomer AL, van Hijum SA, Kuipers OP (2005). "Comparative and functional genomics of lactococci". FEMS Microbiology Reviews. 29 (3): 411–33. doi:10.1016/j.femsre.2005.04.004. PMID 15936843.
- van Hylckama Vlieg JE, Rademaker, JL, Bachmann H, Molenaar D, Kelly WJ, Siezen RJ (2006). "Natural diversity and adaptive responses of Lactococcus lactis". Current Opinion in Biotechnology. 17 (2): 183–90. doi:10.1016/j.copbio.2006.02.007. PMID 16517150.
- Kelly WJ, Ward LJ, Leahy SC (2010). "Chromosomal diversity in Lactococcus lactis and the origin of dairy starter cultures". Genome Biology and Evolution. 2: 729–44. doi:10.1093/gbe/evq056. PMC 2962554. PMID 20847124.
- Passerini D, Beltramo C, Coddeville M, Quentin Y, Ritzenthaler P, Daveran-Mingot ML, Le Bourgeois P (2010). "Genes but Not Genomes Reveal Bacterial Domestication of Lactococcus Lactis". PLOS ONE. 5 (12): e15306. Bibcode:2010PLoSO...515306P. doi:10.1371/journal.pone.0015306. PMC 3003715. PMID 21179431.
- Rademaker JL, Herbet H, Starrenburg MJ, Naser SM, Gevers D, Kelly WJ, Hugenholtz J, et al. (2007). "Diversity analysis of dairy and nondairy Lactococcus lactis isolates, using a novel multilocus sequence analysis scheme and (GTG)5-PCR fingerprinting". Applied and Environmental Microbiology. 73 (22): 7128–37. doi:10.1128/AEM.01017-07. PMC 2168189. PMID 17890345.
- Siezen RJ, Starrenburg MJ, Boekhorst J, Renckens B, Molenaar D, van Hylckama Vlieg JE (2008). "Genome-scale genotype-phenotype matching of two Lactococcus lactis isolates from plants identifies mechanisms of adaptation to the plant niche". Applied and Environmental Microbiology. 74 (2): 424–36. doi:10.1128/AEM.01850-07. PMC 2223259. PMID 18039825.
- Bolotin A, Quinquis B, Renault P, Sorokin A, Ehrlich SD, Kulakauskas S, Lapidus A, et al. (2004). "Complete sequence and comparative genome analysis of the dairy bacterium Streptococcus thermophilus". Nature Biotechnology. 22 (12): 1554–8. doi:10.1038/nbt1034. PMC 7416660. PMID 15543133.
- van de Guchte M, Penaud S, Grimaldi C, Barbe V, Bryson K, Nicolas P, Robert C, et al. (2006). "The complete genome sequence of Lactobacillus bulgaricus reveals extensive and ongoing reductive evolution". Proceedings of the National Academy of Sciences of the United States of America. 103 (24): 9274–9. Bibcode:2006PNAS..103.9274V. doi:10.1073/pnas.0603024103. PMC 1482600. PMID 16754859.
- Bachmann H, Starrenburg MJ, Molenaar D, Kleerebezem M, van Hylckama Vlieg JE (2012). "Microbial domestication signatures of Lactococcus lactis can be reproduced by experimental evolution". Genome Research. 22 (1): 115–24. doi:10.1101/gr.121285.111. PMC 3246198. PMID 22080491.
- Meulen, Sjoerd B. van der; Jong, Anne de; Kok, Jan (2016-03-03). "Transcriptome landscape of Lactococcus lactis reveals many novel RNAs including a small regulatory RNA involved in carbon uptake and metabolism". RNA Biology. 13 (3): 353–366. doi:10.1080/15476286.2016.1146855. ISSN 1547-6286. PMC 4829306. PMID 26950529.
- Schleifer KH, Kraus J, Dvorak C, Kilpper-Bälz R, Collins MD, Fischer W (1985). "Transfer of Streptococcus lactis and Related Streptococci to the Genus Lactococcus gen. nov" (PDF). Systematic and Applied Microbiology. 6 (2): 183–195. doi:10.1016/S0723-2020(85)80052-7. ISSN 0723-2020 – via Elsevier Science Direct.
- Coffey A, Ross RP (2002). "Bacteriophage-resistance systems in dairy starter strains: molecular analysis to application". Antonie van Leeuwenhoek. 82 (1–4): 303–21. doi:10.1023/A:1020639717181. PMID 12369198. S2CID 7217985.
- Davey, Monica (April 15, 2010). "And Now, a State Microbe". New York Times. Retrieved April 19, 2010.
- "No State Microbe For Wisconsin". National Public Radio. Retrieved 28 October 2011.
- "2009 Assembly Bill 556". docs.legis.wisconsin.gov. Retrieved 2017-11-29.
- Madera C, Monjardin C, Suarez JE (2004). "Milk contamination and resistance to processing conditions determine the fate of Lactococcus lactis bacteriophages in dairies". Appl Environ Microbiol. 70 (12): 7365–71. doi:10.1128/AEM.70.12.7365-7371.2004. PMC 535134. PMID 15574937.
- Wyszyńska A, Kobierecka P, Bardowski J, Jagusztyn-Krynicka EK (2015). "Lactic acid bacteria—20 years exploring their potential as live vectors for mucosal vaccination". Appl Microbiol Biotechnol. 99 (7): 2967–2977. doi:10.1007/s00253-015-6498-0. PMC 4365182. PMID 25750046.
- Varma NR, Toosa H, Foo HL, Alitheen NB, Nor Shamsudin M, Arbab AS, Yusoff K, Abdul Rahim R (2013). "Display of the viral epitopes on Lactococcus lactis: a model for food grade vaccine against EV71". Biotechnology Research International. 2013 (11): 4032–4036. doi:10.1155/2013/431315. PMC 431315. PMID 1069289.
- Mierau I, Kleerebezem M (2005). "10 years of the nisin-controlled gene expression system (NICE) in Lactococcus lactis". Appl Microbiol Biotechnol. 68 (6): 705–717. doi:10.1007/s00253-005-0107-6. PMID 16088349. S2CID 24151938.
- Desmond C, Fitzgerald G, Stanton C, Ross R (2004). "Improved stress tolerance of GroESL-overproducing Lactococcus lactis and probiotic Lactobacillus paracasei NFBC 338". Appl Environ Microbiol. 70 (10): 5929–5936. doi:10.1128/AEM.70.10.5929-5936.2004. PMC 522070. PMID 15466535.
- Benbouziane B, Ribelles P, Aubry C, Martin R, Kharrat P, Riazi A, Langella P, Bermudez-Humaran LG (2013). "Development of a Stress-Inducible Controlled Expression (SICE) system in Lactococcus lactis for the production and delivery of therapeutic molecules at mucosal surfaces". J. Biotechnol. 168 (2): 120–129. doi:10.1016/j.jbiotec.2013.04.019. PMID 23664884.
- Nakamura S, Morimoto YV, Kudo S (2015). "A lactose fermentation product produced by Lactococcus lactis subsp. lactis, acetate, inhibits the motility of flagellated pathogenic bacteria". Microbiology. 161 (4): 701–707. doi:10.1099/mic.0.000031. PMID 25573770. S2CID 109572.
- Kihara M, Macnab RM (1981). "Cytoplasmic pH mediates pH taxis and weak-acid repellent taxis of bacteria". J Bacteriol. 145 (3): 1209–1221. doi:10.1128/JB.145.3.1209-1221.1981. PMC 217121. PMID 7009572.
- Repaske DR, Adler J (1981). "Change in intracellular pH of Escherichia coli mediates the chemotactic response to certain attractants and repellents". J Bacteriol. 145 (3): 1196–1208. doi:10.1128/JB.145.3.1196-1208.1981. PMC 217120. PMID 7009571.
- Stordeur P, Goldman M (1998). "Interleukin-10 as a regulatory cytokine induced by cellular stress: molecular aspects". Int. Rev. Immunol. 16 (5–6): 501–522. doi:10.3109/08830189809043006. PMID 9646174.
- Pender SL, et al. (1998). "Suppression of T cell-mediated injury in human gut by interleukin 10: role of matrix metalloproteinases". Gastroenterology. 115 (3): 573–583. doi:10.1016/S0016-5085(98)70136-2. PMID 9721154.
- Steidler L, Hans W, Schotte L, Neirynck S, Obermeier F, Falk W, Fiers W, Remaut E (2000). "Treatment of Murine Colitis by Lactococcus lactis Secreting Interleukin-10". Science. 289 (5483): 1352–1355. Bibcode:2000Sci...289.1352S. doi:10.1126/science.289.5483.1352. PMID 10958782.
- Zhang B, Li A, Zuo F, Yu R, Zeng Z, Ma H, Chen S (2016). "Recombinant Lactococcus lactis NZ9000 secretes a bioactive kisspeptin that inhibits proliferation and migration of human colon carcinoma HT-29 cells". Microbial Cell Factories. 15 (1): 102. doi:10.1186/s12934-016-0506-7. PMC 4901401. PMID 27287327.
- Bauvois B (2012). "New facets of matrix metalloproteinases MMP-2 and MMP-9 as cell surface transducers: outside-in signaling and relationship to tumor progression". Biochim Biophys Acta. 1825 (1): 29–36. doi:10.1016/j.bbcan.2011.10.001. PMID 22020293.
- Kessenbrock K, Plaks V, Werb Z (2010). "Matrix metalloproteinases: regulators of the tumor microenvironment". Cell. 141 (1): 52–67. doi:10.1016/j.cell.2010.03.015. PMC 2862057. PMID 20371345.
- Klein T, Bischoff R (2011). "Physiology and pathophysiology of matrix metalloproteases". Amino Acids. 41 (2): 271–290. doi:10.1007/s00726-010-0689-x. PMC 3102199. PMID 20640864.
- Nash KT, Welch DR (2006). "The KISS1 metastasis suppressor: mechanistic insights and clinical utility". Front Biosci. 11: 647–659. doi:10.2741/1824. PMC 1343480. PMID 16146758.
- Gorbach SL (1990). "Lactic acid bacteria and human health". Ann Med. 22 (1): 37–41. doi:10.3109/07853899009147239. PMID 2109988.
- Hirayama K, Rafter J (1999). "The role of lactic acid bacteria in colon cancer prevention: mechanistic considerations". Antonie van Leeuwenhoek. 76 (1–4): 391–394. doi:10.1007/978-94-017-2027-4_25. ISBN 978-90-481-5312-1. PMID 10532395.
- Ruas-Madiedo P, Hugenholtz J, Zoon P (2002). "An overview of the functionality of exopolysaccharides produced by lactic acid bacteria". Int Dairy J. 12 (2–3): 163–171. doi:10.1016/S0958-6946(01)00160-1.
- Looijesteijn PJ, Trapet L, de Vries E, Abee T, Hugenholtz J (2001). "Physiological function of exopolysaccharides produced by Lactococcus lactis". Int J Food Microbiol. 64 (1–2): 71–80. doi:10.1016/S0168-1605(00)00437-2. PMID 11252513.
- Ji K, Ye L, Ruge F, Hargest R, Mason MD, Jiang WG (2014). "Implication of metastasis suppressor gene, Kiss-1 and its receptor Kiss-1R in colorectal cancer". BMC Cancer. 14: 723. doi:10.1186/1471-2407-14-723. PMC 4190326. PMID 25260785.