Peptidomimetic
A peptidomimetic is a small protein-like chain designed to mimic a peptide.[1] They typically arise either from modification of an existing peptide, or by designing similar systems that mimic peptides, such as peptoids and β-peptides. Irrespective of the approach, the altered chemical structure is designed to advantageously adjust the molecular properties such as stability or biological activity. This can have a role in the development of drug-like compounds from existing peptides. These modifications involve changes to the peptide that will not occur naturally (such as altered backbones and the incorporation of nonnatural amino acids). Based on their similarity with the precursor peptide, peptidomimetics can be grouped into four classes (A – D) where A features the most and D the least similarities. Classes A and B involve peptide-like scaffolds, while classes C and D include small molecules (Figure 1).[2]
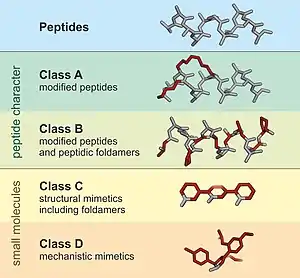
Peptides
D-peptides
A D-peptide is a small sequence of D-amino acids. Since ribosomes are specific to L-amino acids, D-peptides rarely occur naturally in organisms and are not easily digested or degraded. D-peptide peptidomimetics are D-peptides designed to mimic natural L-peptides that commonly have therapeutic properties.
Properties of D-peptides

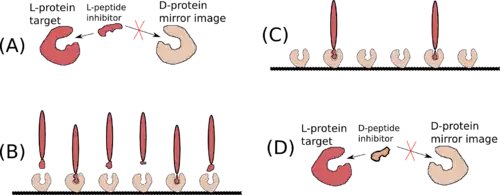
When placed in a nonchiral solvent like water, D-peptides, as well as the larger polypeptide D-proteins, have similar but mirrored properties to the L-peptides and L-proteins with identical sequences. If an L-protein does not require a Chaperone or a structural cofactor to fold, its D-enantiomer protein should have a mirror image conformation with respect to the L-protein (Figure 2). A D-enzyme should act on substrates of reverse chirality compared to the L-enzyme with the same sequence. Similarly, if an L-peptide binds to an L-protein, their D-peptide and D-protein counterparts should bind together in a mirrored way.[3]
D-peptides also have properties that make them attractive as drugs. D-peptides are less susceptible to be degraded in the stomach or inside cells by proteolysis. D-peptide drugs can, therefore, be taken orally and are effective for a longer period of time. D-peptides are easy to synthesize when compared to many other drugs. In some cases, D-peptides can have a low immunogenic response.[4]
Ret design
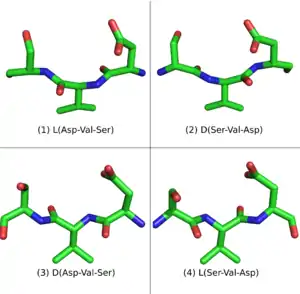
An L-peptide has three analogue sequences (Figure 3) built from L and D amino acids: the D-enantiomer or inverso-peptide with the same sequence, but composed of D-amino acids and a mirror conformation; the retro-peptide, consisting of the same sequence of L amino acids but in reverse order; and the retro-inverso or D-retro-enantiomer peptide, consisting of D-amino acids in the reversed sequence.[5] [6]
While the L-peptide and its D-enantiomer are mirror structures of each other, the L-retro-peptide is the mirror image of the D-retro-inverso-peptide. On the other hand, the L-peptide and the D-retro-inverso-peptide share a similar arrangement of side-chains, although their carboxyl and amino groups point in opposing directions. For small peptides that do not depend on a secondary structure for binding, an L-peptide and its D-retro-inverso-peptide is likely to have a similar binding affinity with a target L-protein.
Mirror-image phage display
Phage display is a technique to screen large libraries of peptides for binding to a target protein. In phage display, the DNA sequence that codes the potential drug-peptide is fused to the gene of the protein coat of bacteriophages and introduced into a vector. Diversity can be introduced to the peptide by mutagenesis. The protein coats peptides are then expressed and purified, and applied to a surface of immobilized protein targets. The surface is then washed away to remove non-binding peptides, while the remaining binding peptides are eluted. [7]
Mirror-image phage display is a similar method that can be used to screen large libraries of D-peptides that bind to target L-proteins. More precisely, since D-peptides can not be expressed in bacteriophages, mirror-image phage display screens L-peptides that bind to immobilized D-proteins that are previously chemically synthesized. Because of the mirror properties of D-peptides, the D-enantiomer of an L-peptide that binds to a D-protein will bind to the L-protein.
Mirror-image phage display, however, has two disadvantages when compared to phage display. Target D-proteins must be chemically synthesized, which is normally an expensive and time-consuming process. Also, the target protein must not require a cofactor or a chaperone to fold, otherwise the chemically synthesized D-protein will not fold to the target, mirror structure.

Structural similarity
Peptide with secondary structure cannot be mimicked by its retro-inverse, as linking in the reverse order breaks many backbone interactions essential for the secondary structure.[8] An approach to mimicking these peptides is by searching for similar (sidechain) structures in a mirrored copy of the Protein Data Bank for the structured elements, and then linking the sections by retro-inversed versions of the loops found in the original protein.[9]
Examples
Peptidomimetic approaches have been utilized to design small molecules that selectively kill cancer cells, an approach known as targeted chemotherapy, by inducing programmed cell death by a process called apoptosis. The following two examples mimic proteins involved in key Protein–protein interactions that reactivate the apoptotic pathway in cancer but do so by distinct mechanisms.
In 2004, Walensky and co-workers reported a stabilized alpha helical peptide that mimics pro-apoptotic BH3-only proteins, such as BID and BAD.[10] This molecule was designed to stabilize the native helical structure by forming a macrocycle between side chains that are not involved in binding. This process, referred to as peptide stapling, uses non-natural amino acids to facilitate macrocyclization by ring-closing olefin metathesis.[11] In this case, a stapled BH3 helix was identified which specifically activates the mitochondrial apoptotic pathway by antagonizing the sequestration of BH3-only proteins by anti-apoptotic proteins (e.g. Bcl-2, see also intrinsic and extrinsic inducers of the apoptosis). This molecule suppressed growth of human leukemia in a mouse xenograft model.[10]
Also in 2004, Harran and co-workers reported a dimeric small molecule that mimics the proapoptotic protein Smac (see mitochondrial regulation in apoptosis).[12] This molecule mimics the N-terminal linear motif Ala-Val-Pro-Ile. Uniquely, the dimeric structure of this peptidomimetic led to a marked increase in activity over an analogous monomer. This binding cooperativity results from the molecule's ability to also mimic the homodimeric structure of Smac, which is functionally important for reactivating caspases.[13] Smac mimetics of this type can sensitize an array of non-small-cell lung cancer cells to conventional chemotherapeutics (e.g. Gemcitabine, Vinorelbine) both in vitro and in mouse xenograft models.[14]
Heterocycles are often used to mimic the amide bond of peptides. Thiazoles, for example, are found in naturally occurring peptides and used by researchers to mimic the amide bond of peptide.[15]
See also
References
- Marshall GR, Ballante F (September 2017). "Limiting Assumptions in the Design of Peptidomimetics". Drug Development Research. 78 (6): 245–267. doi:10.1002/ddr.21406. PMID 28875546. S2CID 5730986.
- Pelay-Gimeno M, Glas A, Koch O, Grossmann TN (July 2015). "Structure-Based Design of Inhibitors of Protein-Protein Interactions: Mimicking Peptide Binding Epitopes". Angewandte Chemie. 54 (31): 8896–927. doi:10.1002/anie.201412070. PMC 4557054. PMID 26119925.
- Milton RC, Milton SC, Kent SB (1992). "Total chemical synthesis of a D-enzyme: the enantiomers of HIV-1 protease show demonstration of reciprocal chiral substrate specificity". Science. 256 (5062): 1445–1448. doi:10.1126/science.1604320. PMID 1604320.
- Welch BD, VanDemark AP, Heroux A, Hill CP, Kay MS (October 2007). "Potent D-peptide inhibitors of HIV-1 entry". Proceedings of the National Academy of Sciences of the United States of America. 104 (43): 16828–33. Bibcode:2007PNAS..10416828W. doi:10.1073/pnas.0708109104. PMC 2040420. PMID 17942675.
- Guichard G, Benkirane N, Zeder-Lutz G, van Regenmortel MH, Briand JP, Muller S (October 1994). "Antigenic mimicry of natural L-peptides with retro-inverso-peptidomimetics". Proceedings of the National Academy of Sciences of the United States of America. 91 (21): 9765–9. Bibcode:1994PNAS...91.9765G. doi:10.1073/pnas.91.21.9765. PMC 44897. PMID 7937888.
- Cardó-Vila M, Giordano RJ, Sidman RL, Bronk LF, Fan Z, Mendelsohn J, Arap W, Pasqualini R (March 2010). "From combinatorial peptide selection to drug prototype (II): targeting the epidermal growth factor receptor pathway". Proceedings of the National Academy of Sciences of the United States of America. 107 (11): 5118–23. Bibcode:2010PNAS..107.5118C. doi:10.1073/pnas.0915146107. PMC 2841862. PMID 20190183.
- Wiesehan K, Willbold D (September 2003). "Mirror-image phage display: aiming at the mirror". ChemBioChem. 4 (9): 811–5. doi:10.1002/cbic.200300570. PMID 12964153. S2CID 34014107.
- Lacroix E, Viguera AR, Serrano L (April 1998). "Reading protein sequences backwards". Folding & Design. 3 (2): 79–85. doi:10.1016/S1359-0278(98)00013-3. PMID 9565752.
- Garton M, Nim S, Stone TA, Wang KE, Deber CM, Kim PM (February 2018). "Method to generate highly stable D-amino acid analogs of bioactive helical peptides using a mirror image of the entire PDB". Proceedings of the National Academy of Sciences of the United States of America. 115 (7): 1505–1510. doi:10.1073/pnas.1711837115. PMC 5816147. PMID 29378946.
- Walensky LD, Kung AL, Escher I, Malia TJ, Barbuto S, Wright RD, Wagner G, Verdine GL, Korsmeyer SJ (September 2004). "Activation of apoptosis in vivo by a hydrocarbon-stapled BH3 helix". Science. 305 (5689): 1466–70. Bibcode:2004Sci...305.1466W. doi:10.1126/science.1099191. PMC 1360987. PMID 15353804.
- Blackwell HE, Grubbs RH (1998). "Highly Efficient Synthesis of Covalently Cross-Linked Peptide Helices by Ring-Closing Metathesis". Angewandte Chemie International Edition. 37 (23): 3281–3284. doi:10.1002/(SICI)1521-3773(19981217)37:23<3281::AID-ANIE3281>3.0.CO;2-V. PMID 29711420.
- Li L, Thomas RM, Suzuki H, De Brabander JK, Wang X, Harran PG (September 2004). "A small molecule Smac mimic potentiates TRAIL- and TNFalpha-mediated cell death". Science. 305 (5689): 1471–4. Bibcode:2004Sci...305.1471L. doi:10.1126/science.1098231. PMID 15353805. S2CID 58926089.
- Chai J, Du C, Wu JW, Kyin S, Wang X, Shi Y (August 2000). "Structural and biochemical basis of apoptotic activation by Smac/DIABLO". Nature. 406 (6798): 855–62. Bibcode:2000Natur.406..855C. doi:10.1038/35022514. PMID 10972280. S2CID 4385614.
- Greer RM, Peyton M, Larsen JE, Girard L, Xie Y, Gazdar AF, Harran P, Wang L, Brekken RA, Wang X, Minna JD (December 2011). "SMAC mimetic (JP1201) sensitizes non-small cell lung cancers to multiple chemotherapy agents in an IAP-dependent but TNF-α-independent manner". Cancer Research. 71 (24): 7640–8. doi:10.1158/0008-5472.CAN-10-3947. PMC 3382117. PMID 22049529.
- Mak JY, Xu W, Fairlie DP (2015-01-01). Peptidomimetics I (PDF). Topics in Heterocyclic Chemistry. 48. Springer Berlin Heidelberg. pp. 235–266. doi:10.1007/7081_2015_176. ISBN 978-3-319-49117-2.
Further reading
- Denicourt C, Dowdy SF (September 2004). "Medicine. Targeting apoptotic pathways in cancer cells". Science. 305 (5689): 1411–3. doi:10.1126/science.1102974. PMID 15353788. S2CID 83339121.