Bradford protein assay
The Bradford protein assay was developed by Marion M. Bradford in 1976.[1] It is a quick and accurate[2] spectroscopic analytical procedure used to measure the concentration of protein in a solution. The reaction is dependent on the amino acid composition of the measured proteins.
Principle
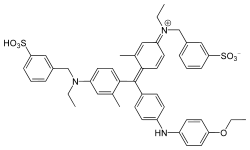

The Bradford assay, a colorimetric protein assay, is based on an absorbance shift of the dye Coomassie Brilliant Blue G-250. The Coomassie Brilliant Blue G-250 dye exists in three forms: anionic (blue), neutral (green), and cationic (red).[3] Under acidic conditions, the red form of the dye is converted into its blue form, binding to the protein being assayed. If there's no protein to bind, then the solution will remain brown. The dye forms a strong, noncovalent complex with the protein's carboxyl group by van der Waals force and amino group through electrostatic interactions.[1] During the formation of this complex, the red form of Coomassie dye first donates its free electron to the ionizable groups on the protein, which causes a disruption of the protein's native state, consequently exposing its hydrophobic pockets. These pockets in the protein's tertiary structure bind non-covalently to the non-polar region of the dye via the first bond interaction (van der Waals forces) which position the positive amine groups in proximity with the negative charge of the dye. The bond is further strengthened by the second bond interaction between the two, the ionic interaction. When the dye binds to the protein, it causes a shift from 465 nm to 595 nm , which is why the absorbance readings are taken at 595 nm.[4]
The cationic (unbound) form is green / red and has an absorption spectrum maximum historically held to be at 465 nm. The anionic bound form of the dye which is held together by hydrophobic and ionic interactions, has an absorption spectrum maximum historically held to be at 595 nm.[5] The increase of absorbance at 595 nm is proportional to the amount of bound dye, and thus to the amount (concentration) of protein present in the sample.
Unlike other protein assays, the Bradford protein assay is less susceptible to interference by various chemical compounds such as sodium, potassium or even carbohydrates like sucrose, that may be present in protein samples.[2] An exception of note is elevated concentrations of detergent. Sodium dodecyl sulfate (SDS), a common detergent, may be found in protein extracts because it is used to lyse cells by disrupting the membrane lipid bilayer and to denature proteins for SDS-PAGE. While other detergents interfere with the assay at high concentration, the interference caused by SDS is of two different modes, and each occurs at a different concentration. When SDS concentrations are below critical micelle concentration (known as CMC, 0.00333%W/V to 0.0667%) in a Coomassie dye solution, the detergent tends to bind strongly with the protein, inhibiting the protein binding sites for the dye reagent. This can cause underestimations of protein concentration in solution. When SDS concentrations are above CMC, the detergent associates strongly with the green form of the Coomassie dye, causing the equilibrium to shift, thereby producing more of the blue form. This causes an increase in the absorbance at 595 nm independent of protein presence.
Other interference may come from the buffer used when preparing the protein sample. A high concentration of buffer will cause an overestimated protein concentration due to depletion of free protons from the solution by conjugate base from the buffer. This will not be a problem if a low concentration of protein (subsequently the buffer) is used.
In order to measure the absorbance of a colorless compound a Bradford assay must be performed. Some colorless compounds such as proteins can be quantified at an Optical Density of 280 nm due to the presence of aromatic rings such as Tryptophan, Tyrosine and Phenylalanine but if none of these amino acids are present then the absorption cannot be measured at 280 nm.[6]
Advantages
Many protein-containing solutions have the highest absorption at 280 nm in the spectrophotometer, the UV range. This requires spectrophotometers capable of measuring in the UV range, which many cannot. Additionally, the absorption maxima at 280 nm requires that proteins contain aromatic amino acids such as tyrosine (Y), phenylalanine (F) and/or tryptophan (W). Not all proteins contain these amino acids, a fact which will skew the concentration measurements. If nucleic acids are present in the sample, they would also absorb light at 280 nm, skewing the results further. By using the Bradford protein assay, one can avoid all of these complications by simply mixing the protein samples with the Coomassie Brilliant Blue G-250 dye (Bradford reagent) and measuring their absorbances at 595 nm, which is in the visible range.[7]
The procedure for Bradford protein assay is very easy and simple to follow. It is done in one step where the Bradford reagent is added to a test tube along with the sample. After mixing well, the mixture almost immediately changes to a blue color. When the dye binds to the proteins through a process that takes about 2 minutes, a change in the absorption maximum of the dye from 465 nm to 595 nm in acidic solutions occurs.[2] This dye creates strong noncovalent bonds with the proteins, via electrostatic interactions with the amino and carboxyl groups, as well as Van Der Waals interactions. Only the molecules that bind to the proteins in solution exhibit this change in absorption, which eliminates the concern that unbound molecules of the dye might contribute to the experimentally obtained absorption reading. This process is more beneficial since it is less pricey than other methods, easy to use, and has high sensitivity of the dye for protein.[8]
After 5 minutes of incubation, the absorbance can be read at 595 nm using a spectrophotometer; an easily accessible machine.
This assay is one of the fastest assays performed on proteins.[9] The total time it takes to set up and complete the assay is under 30 minutes.[10] The entire experiment is done at room temperature.
The Bradford protein assay can measure protein quantities as little as 1 to 20 μg.[11] It is an extremely sensitive technique.
The dye reagent is a stable ready to use product prepared in phosphoric acid. It can remain at room temperature for up to 2 weeks before it starts to degrade.
Protein samples usually contain salts, solvents, buffers, preservatives, reducing agents and metal chelating agents. These molecules are frequently used for solubilizing and stabilizing proteins. Other protein assay like BCA and Lowry are ineffective because molecules like reducing agents interfere with the assay.[12] Using Bradford can be advantageous against these molecules because they are compatible to each other and will not interfere.[13]
The linear graph acquired from the assay (absorbance versus protein concentration in μg/mL) can be easily extrapolated to determine the concentration of proteins by using the slope of the line.
It is a sensitive technique. It is also very simple: measuring the OD at 595 nm after 5 minutes of incubation. This method can also make use of a Vis spectrophotometer.[14]
Disadvantages
The Bradford assay is linear over a short range, typically from 0 µg/mL to 2000 µg/mL, often making dilutions of a sample necessary before analysis. In making these dilutions, error in one dilution is compounded in further dilutions resulting in a linear relationship that may not always be accurate.
Basic conditions and detergents, such as SDS, can interfere with the dye's ability to bind to the protein through its side chains.[9] However, there are some detergent-compatible Bradford reagents. The Bradford assay depends on the sequence of the protein. Thus, if the protein does not contain an ideal number of aromatic residues, then the dye will not be able to bind to the protein efficiently. Another disadvantage of the Bradford Protein Assay is that this method depends on comparing the absorbance of the protein to that of a standard protein. If the protein does not react to the dye in a similar way as the standard protein, it is possible that the concentration measured will be inaccurate.
The reagents in this method tend to stain the test tubes. Same test tubes cannot be used since the stain would affect the absorbance reading. This method is also time sensitive. When more than one solution is tested, it is important to make sure every sample is incubated for the same amount of time for accurate comparison.[15]
It is also inhibited by the presence of detergents, although this problem can be alleviated by the addition of cyclodextrins to the assay mixture.[16]
Much of the non-linearity stems from the equilibrium between two different forms of the dye which is perturbed by adding the protein. The Bradford assay linearizes by measuring the ratio of the absorbances, 595 over 450 nm. This modified Bradford assay is approximately 10 times more sensitive than the conventional one.[17]
The Coomassie Blue G250 dye used to bind to the proteins in the original Bradford method readily binds to arginine and lysine groups of proteins. This is a disadvantage because the preference of the dye to bind to these amino acids can result in a varied response of the assay between different proteins. Changes to the original method, such as increasing the pH by adding NaOH or adding more dye have been made to correct this variation. Although these modifications result in a less sensitive assay, a modified method becomes sensitive to detergents that can interfere with sample.[18]
Sample Bradford procedure
Materials
- Lyophilized bovine plasma gamma globulin
- Coomassie Brilliant Blue 1
- 0.15 M NaCl
- Spectrophotometer and cuvettes
- Micropipettes
Procedure (Standard Assay, 20-150 µg protein; 200-1500 µg/mL)
- Prepare a series of standards diluted with 0.15 M NaCl to final concentrations of 0 (blank = No protein), 250, 500, 750 and 1500 µg/mL. Also prepare serial dilutions of the unknown sample to be measured.
- Add 100 µL of each of the above to a separate test tube (or spectrophotometer tube if using a Spectronic 20).
- Add 5.0 mL of Coomassie Blue to each tube and mix by vortex, or inversion.
- Adjust the spectrophotometer to a wavelength of 595 nm, using the tube which contains no protein (blank).
- Wait 5 minutes and read each of the standards and each of the samples at 595 nm wavelength.
- Plot the absorbance of the standards vs. their concentration. Compute the extinction coefficient and calculate the concentrations of the unknown samples.
Procedure (Micro Assay, 1-10 µg protein/mL)
- Prepare standard concentrations of protein of 1, 5, 7.5 and 10 µg/mL. Prepare a blank of NaCl only. Prepare a series of sample dilutions.
- Add 100 µL of each of the above to separate tubes (use microcentrifuge tubes) and add 1.0 mL of Coomassie Blue to each tube.
- Turn on and adjust a spectrophotometer to a wavelength of 595 nm, and blank the spectrophotometer using 1.5 mL cuvettes.
- Wait 2 minutes and read the absorbance of each standard and sample at 595 nm.
- Plot the absorbance of the standards vs. their concentration. Compute the extinction coefficient and calculate the concentrations of the unknown samples.
Using data obtained to find concentration of unknown
In summary, in order to find a standard curve, one must use varying concentrations of BSA (Bovine Serum Albumin)[2] in order to create a standard curve with concentration plotted on the x-axis and absorbance plotted on the y-axis. Only a narrow concentration of BSA is used (2-10 ug/mL) in order to create an accurate standard curve.[19] Using a broad range of protein concentration will make it harder to determine the concentration of the unknown protein. This standard curve is then used to determine the concentration of the unknown protein. The following elaborates on how one goes from the standard curve to the concentration of the unknown.
First, add a line of best fit, or Linear regression and display the equation on the chart. Ideally, the R2 value will be as close to 1 as possible. R represents the sum of the square values of the fit subtracted from each data point. Therefore, if R2 is much less than one, consider redoing the experiment to get one with more reliable data.[20]
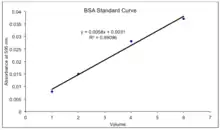
The equation displayed on the chart gives a means for calculating the absorbance and therefore concentration of the unknown samples. In Graph 1, x is concentration and y is absorbance, so one must rearrange the equation to solve for x and enter the absorbance of the measured unknown.[21] It is likely that the unknown will have absorbance numbers outside the range of the standard. These should not be included calculations, as the equation given cannot apply to numbers outside of its limitations. In a large scale, one must compute the extinction coefficient using the Beer-Lambert Law A=εLC in which A is the measured absorbance, ε is the slope of the standard curve, L is the length of the cuvette, and C is the concentration being determined.[22] In a micro scale, a cuvette may not be used and therefore one only has to rearrange to solve for x.

In order to attain a concentration that makes sense with the data, the dilutions, concentrations, and units of the unknown must be normalized (Table 1). To do this, one must divide concentration by volume of protein in order to normalize concentration and multiply by amount diluted to correct for any dilution made in the protein before performing the assay.
Alternative assays
Alternative protein assays include:
References
- Ninfa, Alexander J; Ballou, David P; Benore, Marilee (2008). Fundamental Laboratory Approaches for Biochemistry and Biotechnology. Wiley. p. 113.
- Bradford, Marion (1976). "A Rapid and Sensitive Method for the Quantification of Microgram Quantities of Protein Utilizing the Principle of Protein-Dye Binding" (PDF). Analytical Biochemistry. 72 (1–2): 248–254. doi:10.1006/abio.1976.9999. PMID 942051 – via Google Scholar.
- "Quick Start TM Bradford Protein Assay" (PDF). www.bio-rad.com.
- Bradford, Marion (1976). "A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding". Analytical Biochemistry. 72 (1–2): 248–254. doi:10.1006/abio.1976.9999. PMID 942051 – via ScienceDirect.
- "Protein determination by the Bradford method".
- P., Ballou, David; Marilee., Benore (2010). Fundamental laboratory approaches for biochemistry and biotechnology. John Wiley. ISBN 9780470087664. OCLC 420027217.
- Ninfa, Ballou, Benore, Alexander J., David P., Marilee (2010). Fundamental Laboratory Approaches for Biochemistry and Biotechnology. United States of America: John Wiley & Sons, Inc. pp. 110, 113. ISBN 978-0-470-08766-4.CS1 maint: multiple names: authors list (link)
- Ninfa, Alexander J.; Ballou, David P.; Benore, Marilee (2010). Fundamental Laboratory Approaches for Biochemistry and Biotechnology. John Wiley & Sons Inc. p. 113. ISBN 978-0470087664.
- Okutucu, Burcu; Dınçer, Ayşşe; Habib, Ömer; Zıhnıoglu, Figen (2007-08-01). "Comparison of five methods for determination of total plasma protein concentration". Journal of Biochemical and Biophysical Methods. 70 (5): 709–711. doi:10.1016/j.jbbm.2007.05.009. PMID 17597224.
- "Protein Assay Technical Handbook" (PDF).
- "4.5. Determination of protein concentration". elte.prompt.hu. Archived from the original on 2016-09-21. Retrieved 2016-05-19.
- barbosa, Helder; Slater K.H., Nigel (3 August 2009). "Protein quantification in the presence of poly(ethylene glycol) and dextran using the Bradford method". Analytical Biochemistry: Methods in the Biological Sciences. 395 (1): 108–110. doi:10.1016/j.ab.2009.07.045. PMID 19653991.
- Ninfa, Alexander J. (2010). Fundamental Laboratory Approaches for Biochemistry and Biotechnology. Wiley. pp. 117–118. ISBN 978-0470087664.
- Ninfa, Ballou (1998). Fundamental Approaches to Biochemistry and Biotechnology. Fitzgerald Science Press, Bethesda, MD. pp. 114–116. ISBN 978-0470087664.
- Ninfa, Alexander J; Ballou, David P; Benore, Marilee (2009). Fundamental Laboratory Approaches for Biochemistry and Biotechnology. Wiley. p. 113.
- Rabilloud, Thierry (2018). "Optimization of the cydex blue assay: A one-step colorimetric protein assay using cyclodextrins and compatible with detergents and reducers". PLOS ONE. 13 (4): e0195755. Bibcode:2018PLoSO..1395755R. doi:10.1371/journal.pone.0195755. PMC 5895047. PMID 29641569.
- Zor, Tsaffrir; Selinger, Zvi (1996-05-01). "Linearization of the Bradford Protein Assay Increases Its Sensitivity: Theoretical and Experimental Studies". Analytical Biochemistry. 236 (2): 302–308. doi:10.1006/abio.1996.0171. PMID 8660509.
- Kruger (2002). "The Bradford Method for Protein Quantitation". The Bradford Method for Protein Quantification. pp. 15–22. doi:10.1385/1-59259-169-8:15. ISBN 1-59259-169-8. S2CID 36834925.
- "Linearization of the Bradford Protein Assay Increases Its Sensitivity: Theoretical and Experimental Studies" (PDF). www.tau.ac. November 20, 1995.
- Albright, Brian (2009). Mathematical Modeling with Excel. p. 60. ISBN 978-0763765668.
- Stephenson, Frank Harold (2003). Calculations for molecular biology and biotechnology: a guide to mathematics in the laboratory. pp. 252. ISBN 978-0126657517.
- Ibanez, Jorge G. (2007). Environmental chemistry: fundamentals. pp. 60. ISBN 978-0387260617.
Further reading
- Bradford, M.M. (1976), "Rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding", Anal. Biochem., 72 (1–2): 248–254, doi:10.1016/0003-2697(76)90527-3, PMID 942051
- Zor, T.; Selinger, Z. (1996), "Linearization of the Bradford protein assay increases its sensitivity: theoretical and experimental studies", Anal. Biochem., 236 (2): 302–308, doi:10.1006/abio.1996.0171, PMID 8660509
- Noble, J.E.; Bailey, M.J.A. (2009), "Quantitation of Protein", Methods Enzymol., Methods in Enzymology, 463: 73–95, doi:10.1016/S0076-6879(09)63008-1, ISBN 9780123745361, PMID 19892168
- Albright, Brian (2009), Mathematical Modeling with Excel, p. 60, ISBN 978-0763765668
- Stephenson, Frank Harold (2003), Calculations for molecular biology and biotechnology: a guide to mathematics in the laboratory, pp. 252, ISBN 978-0126657517
- Dennison, Clive (2003), "A guide to protein isolation", Focus on Structural Biology, 3: 39, ISBN 978-1402012242
- Ibanez, Jorge G. (2007), Environmental chemistry: fundamentals, p. 60, ISBN 978-0387260617