Small-angle neutron scattering
Small-angle neutron scattering (SANS) is an experimental technique that uses elastic neutron scattering at small scattering angles to investigate the structure of various substances at a mesoscopic scale of about 1–100 nm.
| Science with neutrons |
|---|
 |
| Foundations |
| Neutron scattering |
| Other applications |
|
| Infrastructure |
|
| Neutron facilities |
Small angle neutron scattering is in many respects very similar to small-angle X-ray scattering (SAXS); both techniques are jointly referred to as small-angle scattering (SAS). Advantages of SANS over SAXS are its sensitivity to light elements, the possibility of isotope labelling, and the strong scattering by magnetic moments.
Technique
During a SANS experiment a beam of neutrons is directed at a sample, which can be an aqueous solution, a solid, a powder, or a crystal. The neutrons are elastically scattered by nuclear interaction with the nuclei or interaction with magnetic momentum of unpaired electrons. In X-ray scattering, photons interact with the electronic cloud so the bigger the element, the bigger the effect is. In neutron scattering, neutrons interact with nuclei and the interaction depends on the isotope; some light elements like deuterium show similar scattering cross section as heavy elements like Pb.
In zero order dynamical theory of diffraction the refractive index is directly related to the scattering length density and is a measure of the strength of the interaction of a neutron wave with a given nucleus. The following table shows the neutron scattering length for a few chemical elements (in 10−12 cm).[1]
| H | D | C | N | O | P | S |
|---|---|---|---|---|---|---|
| −0.3742 | 0.6671 | 0.6651 | 0.940 | 0.5804 | 0.517 | 0.2847 |
Note that the relative scale of the scattering lengths is the same. Another important point is that the scattering from hydrogen is distinct from that of deuterium. Also, hydrogen is one of the few elements that has a negative scatter, which means that neutrons deflected from hydrogen are 180° out of phase relative to those deflected by the other elements. These features are important for the technique of contrast variation (see below).
Related techniques
SANS usually uses collimation of the neutron beam to determine the scattering angle of a neutron, which results in an ever lower signal-to-noise ratio for data that contains information on the properties of a sample at relatively long length scales, beyond ~1 μm. The traditional solution is to increase the brightness of the source, as in Ultra Small Angle Neutron Scattering (USANS). As an alternative Spin-echo Small-angle Neutron Scattering (SESANS) was introduced, using neutron spin echo to track the scattering angle, and expanding the range of length scales which can be studied by neutron scattering to well beyond 10 μm.
Grazing-incidence small-angle scattering (GISANS) combines ideas of SANS and of neutron reflectometry.
In biology
A crucial feature of SANS that makes it particularly useful for the biological sciences is the special behavior of hydrogen, especially compared to deuterium. In biological systems hydrogen can be exchanged with deuterium which usually has minimal effect on the sample but has dramatic effects on the scattering.
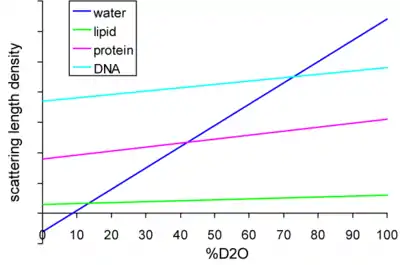
The technique of contrast variation (or contrast matching) relies on the differential scatter of hydrogen vs. deuterium. Figure 1 shows the scattering length density for water and various biological macromolecules as a function of the deuterium concentration. (Adapted from.[1]) Biological samples are usually dissolved in water, so their hydrogens are able to exchange with any deuteriums in the solvent. Since the overall scatter of a molecule depends on the scatter of all its components, this will depend on the ratio of hydrogen to deuterium in the molecule. At certain ratios of H2O to D2O, called match points, the scatter from the molecule will equal that of the solvent, and thus be eliminated when the scatter from the buffer is subtracted from the data. For instance the match point for proteins is typically around 40–45% D2O, and at that concentration the scatter from the protein will be indistinguishable from that of the buffer.
To use contrast variation, different components of a system must scatter differently. This can be based on inherent scattering differences, e.g. DNA vs. protein, or arise from differentially labeled components, e.g. having one protein in a complex deuterated while the rest are protonated. In terms of modelling, small-angle X-ray and neutron scattering data can be combined with the program MONSA. An example in which SAXS, SANS and EM data has been used to build an atomic model of a large multi-subunit enzyme has recently been published.[2] For some examples of this method see.[3]
Instruments
There are numerous SANS instruments available worldwide at neutron facilities such as research reactors or spallation sources.
See also
References
- Jacrot, B (1976). "The study of biological structures by neutron scattering from solution". Reports on Progress in Physics. 39 (10): 911–53. Bibcode:1976RPPh...39..911J. doi:10.1088/0034-4885/39/10/001.
- Kennaway, Chris; Taylor, James; et al. (1 Jan 2012). "Structure and operation of the DNA-translocating type I DNA restriction enzymes". Genes & Development. 26 (4): 92–104. doi:10.1101/gad.179085.111. PMC 3258970. PMID 22215814.
- Perkins, SJ (January 1, 1988). "Structural studies of proteins by high-flux X-ray and neutron solution scattering". Biochemical Journal. 254 (2): 313–27. PMC 1135080. PMID 3052433.
Textbooks
- Fejgin, Lev A.: Structure analysis by small-angle X-ray and neutron scattering. New York: Plenum (1987).
- Higgins, Julia S.; Benoît, Henri: Polymers and neutron scattering. Oxford: Clarendon Press (1994?).
External links
- The Small-Angle Scattering portal, link collection, with elaborate software list
- World directory of SANS instruments
- B. Hammouda: Probing Nanoscale Structures – The SANS Toolbox (690 pages)
- Small Angle Scattering at ISIS Neutron and Muon Source
