Protein tandem repeats
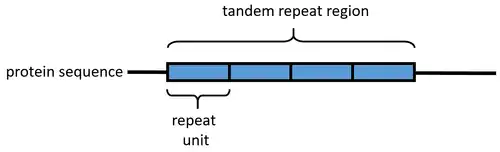
An array of protein tandem repeats is defined as several (at least two) adjacent copies having the same or similar sequence motifs. These periodic sequences are generated by internal duplications in both coding and non-coding genomic sequences. Repetitive units of protein tandem repeats are considerably diverse, ranging from the repetition of a single amino acid to domains of 100 or more residues.[1][2]
"Repeats" in proteins

In proteins, a "repeat" is any sequence block that returns more than one time in the sequence, either in an identical or a highly similar form. The degree of similarity can be highly variable, with some repeats maintaining only a few conserved amino acid positions and a characteristic length. Highly degenerate repeats can be very difficult to detect from sequence alone. Structural similarity can help to identify repetitive patterns in sequence.
Structure
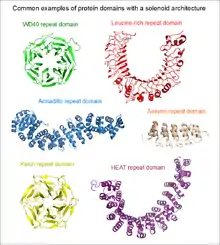
Repetitiveness does not in itself indicate anything about the structure of the protein. As a "rule of thumb", short repetitive sequences (e.g. those below the length of 10 amino acids) may be intrinsically disordered, and not part of any folded protein domains. Repeats that are at least 30 to 40 amino acids long, are far more likely to be folded as part of a domain. Such long repeats are frequently indicative of the presence of a solenoid domain in the protein.
Approximately half of the tandem repeat regions have intrinsically disordered conformation being naturally unfolded.[3][4][5] Examples of disordered repetitive sequences include the 7-mer peptide repeats found in the RPB1 subunit of RNA polymerase II,[6] or the tandem beta-catenin or axin binding linear motifs in APC (adenomatous polyposis coli).[7] The other half of the regions with the stable 3D structure has a plethora of shapes and functions.[8][9] Examples of short repeats exhibiting ordered structures include the three-residue collagen repeat or the five-residue pentapeptide repeat that forms a beta helix structure.
Classification
Depending on the length of the repetitive units, their protein structures can be subdivided into five classes:[8][9]
- crystalline aggregates formed by regions with 1 or 2 residue long repeats, archetypical low complexity regions
- fibrous structures stabilized by inter-chain interactions with 3-7 residue repeats
- elongated structures with repeats of 5–40 residues dominated by solenoid proteins
- closed (not elongated) structures with repeats of 30-60 residues as toroid repeats
- beads on a string structures with typical size of repeats over 50 residues, which are already large enough to fold independently into stable domains.
Function
Some well-known examples of proteins with tandem repeats are collagen, which plays a key role in the arrangement of the extracellular matrix; alpha-helical coiled coils having structural and oligomerization functions; leucine-rich repeat proteins, which specifically bind a number of globular proteins by their concave surfaces; and zinc-finger proteins, which regulate the expression of genes by binding DNA.
Tandem repeat proteins frequently function as protein-protein interaction modules. The WD40 repeat is a prime example of this function.[10]
Distribution in proteomes
Tandem repeats are ubiquitous in proteomes and occur in at least 14% of all proteins.[11] For example, they are present in almost every third human protein and even in every second protein from Plasmodium falciparum or Dictyostelium discoideum.[11][12] Tandem repeats with short repetitive units (especially homorepeats) are more frequent than others.[11]
Annotation methods
Protein tandem repeats can be either detected from sequence or annotated from structure. Specialized methods were built for the identification of repeat proteins.[13]
Sequence-based strategies, based on homology search [14] or domain assignment [15] [16],mostly underestimate TRs due to the presence of highly degenerate repeat units.[17] A recent study to understand and improve Pfam coverage of the human proteome [17] showed that five among the ten largest sequence clusters not annotated with Pfam are repeat regions. Alternatively, methods requiring no prior knowledge for the detection of repeated substrings can be based on self-comparison,[18][19] clustering [20] [21] or hidden Markov models.[22][23] Some others rely on complexity measurements [13] or take advantage of meta searches to combine outputs from different sources.[24][25]
Structure-based methods instead take advantage of the modularity of available PDB structures to recognize repetitive elements.[26][27][28][29][30]
References
- Heringa J (June 1998). "Detection of internal repeats: how common are they?". Current Opinion in Structural Biology. 8 (3): 338–45. doi:10.1016/s0959-440x(98)80068-7. PMID 9666330.
- Andrade MA, Ponting CP, Gibson TJ, Bork P (May 2000). "Homology-based method for identification of protein repeats using statistical significance estimates". Journal of Molecular Biology. 298 (3): 521–37. doi:10.1006/jmbi.2000.3684. PMID 10772867.
- Tompa P (September 2003). "Intrinsically unstructured proteins evolve by repeat expansion". BioEssays. 25 (9): 847–55. doi:10.1002/bies.10324. PMID 12938174. S2CID 32684524.
- Simon M, Hancock JM (2009). "Tandem and cryptic amino acid repeats accumulate in disordered regions of proteins". Genome Biology. 10 (6): R59. doi:10.1186/gb-2009-10-6-r59. PMC 2718493. PMID 19486509.
- Jorda J, Xue B, Uversky VN, Kajava AV (June 2010). "Protein tandem repeats - the more perfect, the less structured" (PDF). The FEBS Journal. 277 (12): 2673–82. doi:10.1111/j.1742-4658.2010.07684.x. PMC 2928880. PMID 20553501.
- Meyer PA, Ye P, Zhang M, Suh MH, Fu J (June 2006). "Phasing RNA polymerase II using intrinsically bound Zn atoms: an updated structural model". Structure. 14 (6): 973–82. doi:10.1016/j.str.2006.04.003. PMID 16765890.
- Liu J, Xing Y, Hinds TR, Zheng J, Xu W (June 2006). "The third 20 amino acid repeat is the tightest binding site of APC for beta-catenin". J. Mol. Biol. 360 (1): 133–44. doi:10.1016/j.jmb.2006.04.064. PMID 16753179.
- Kajava AV (September 2012). "Tandem repeats in proteins: from sequence to structure". Journal of Structural Biology. 179 (3): 279–88. doi:10.1016/j.jsb.2011.08.009. PMID 21884799.
- Paladin L, Hirsh L, Piovesan D, Andrade-Navarro MA, Kajava AV, Tosatto SC (January 2017). "RepeatsDB 2.0: improved annotation, classification, search and visualization of repeat protein structures". Nucleic Acids Research. 45 (D1): D308–D312. doi:10.1093/nar/gkw1136. PMC 5210593. PMID 27899671.
- Stirnimann CU, Petsalaki E, Russell RB, Müller CW (October 2010). "WD40 proteins propel cellular networks". Trends in Biochemical Sciences. 35 (10): 565–74. doi:10.1016/j.tibs.2010.04.003. PMID 20451393.
- Marcotte EM, Pellegrini M, Yeates TO, Eisenberg D (October 1999). "A census of protein repeats". Journal of Molecular Biology. 293 (1): 151–60. doi:10.1006/jmbi.1999.3136. PMID 10512723.
- Pellegrini M (2015). "Tandem Repeats in Proteins: Prediction Algorithms and Biological Role". Frontiers in Bioengineering and Biotechnology. 3: 143. doi:10.3389/fbioe.2015.00143. PMC 4585158. PMID 26442257.
- Pellegrini M, Renda ME, Vecchio A (2012). "Ab initio detection of fuzzy amino acid tandem repeats in protein sequences". BMC Bioinformatics. 13 Suppl 3: S8. doi:10.1186/1471-2105-13-S3-S8. PMC 3402919. PMID 22536906.CS1 maint: multiple names: authors list (link)
- Andrade MA, Ponting CP, Gibson TJ, Bork P (2000). "Homology-based method for identification of protein repeats using statistical significance estimates". J Mol Biol. 298 (3): 521–37. doi:10.1006/jmbi.2000.3684. PMID 10772867.CS1 maint: multiple names: authors list (link)
- El-Gebali S, Mistry J, Bateman A, Eddy SR, Luciani A, Potter SC; et al. (2019). "The Pfam protein families database in 2019". Nucleic Acids Res. 47 (D1): D427–D432. doi:10.1093/nar/gky995. PMC 6324024. PMID 30357350.CS1 maint: multiple names: authors list (link)
- Mitchell AL, Attwood TK, Babbitt PC, Blum M, Bork P, Bridge A; et al. (2019). "InterPro in 2019: improving coverage, classification and access to protein sequence annotations". Nucleic Acids Res. 47 (D1): D351–D360. doi:10.1093/nar/gky1100. PMC 6323941. PMID 30398656.CS1 maint: multiple names: authors list (link)
- Mistry J, Coggill P, Eberhardt RY, Deiana A, Giansanti A, Finn RD; et al. (2013). "The challenge of increasing Pfam coverage of the human proteome". Database (Oxford). 2013: bat023. doi:10.1093/database/bat023. PMC 3630804. PMID 23603847.CS1 maint: multiple names: authors list (link)
- Heger A, Holm L (2000). "Rapid automatic detection and alignment of repeats in protein sequences". Proteins. 41 (2): 224–37. doi:10.1002/1097-0134(20001101)41:2<224::aid-prot70>3.0.co;2-z. PMID 10966575.
- Szklarczyk R, Heringa J (2004). "Tracking repeats using significance and transitivity". Bioinformatics. 20 Suppl 1: i311-7. doi:10.1093/bioinformatics/bth911. PMID 15262814.
- Newman AM, Cooper JB (2007). "XSTREAM: a practical algorithm for identification and architecture modeling of tandem repeats in protein sequences". BMC Bioinformatics. 8: 382. doi:10.1186/1471-2105-8-382. PMC 2233649. PMID 17931424.
- Jorda J, Kajava AV (2009). "T-REKS: identification of Tandem REpeats in sequences with a K-meanS based algorithm". Bioinformatics. 25 (20): 2632–8. doi:10.1093/bioinformatics/btp482. PMID 19671691.
- Söding J, Remmert M, Biegert A (2006). "HHrep: de novo protein repeat detection and the origin of TIM barrels". Nucleic Acids Res. 34 (Web Server issue): W137-42. doi:10.1093/nar/gkl130. PMC 1538828. PMID 16844977.CS1 maint: multiple names: authors list (link)
- Biegert A, Söding J (2008). "De novo identification of highly diverged protein repeats by probabilistic consistency". Bioinformatics. 24 (6): 807–14. doi:10.1093/bioinformatics/btn039. PMID 18245125.
- Gruber M, Söding J, Lupas AN (2005). "REPPER--repeats and their periodicities in fibrous proteins". Nucleic Acids Res. 33 (Web Server issue): W239-43. doi:10.1093/nar/gki405. PMC 1160166. PMID 15980460.CS1 maint: multiple names: authors list (link)
- Schaper E, Anisimova M (2015). "The evolution and function of protein tandem repeats in plants". New Phytol. 206 (1): 397–410. doi:10.1111/nph.13184. PMID 25420631.
- Abraham AL, Rocha EP, Pothier J (2008). "Swelfe: a detector of internal repeats in sequences and structures". Bioinformatics. 24 (13): 1536–7. doi:10.1093/bioinformatics/btn234. PMC 2718673. PMID 18487242.CS1 maint: multiple names: authors list (link)
- Sabarinathan R, Basu R, Sekar K (2010). "ProSTRIP: A method to find similar structural repeats in three-dimensional protein structures". Comput Biol Chem. 34 (2): 126–30. doi:10.1016/j.compbiolchem.2010.03.006. PMID 20430700.CS1 maint: multiple names: authors list (link)
- Walsh I, Sirocco FG, Minervini G, Di Domenico T, Ferrari C, Tosatto SC (2012). "RAPHAEL: recognition, periodicity and insertion assignment of solenoid protein structures". Bioinformatics. 28 (24): 3257–64. doi:10.1093/bioinformatics/bts550. PMID 22962341.CS1 maint: multiple names: authors list (link)
- Hrabe T, Godzik A (2014). "ConSole: using modularity of contact maps to locate solenoid domains in protein structures". BMC Bioinformatics. 15: 119. doi:10.1186/1471-2105-15-119. PMC 4021314. PMID 24766872.
- Do Viet P, Roche DB, Kajava AV (2015). "TAPO: A combined method for the identification of tandem repeats in protein structures". FEBS Lett. 589 (19 Pt A): 2611–9. doi:10.1016/j.febslet.2015.08.025. PMID 26320412. S2CID 28423787.CS1 maint: multiple names: authors list (link)