Retrotransposon
Retrotransposons (also called Class I transposable elements or transposons via RNA intermediates) are a type of genetic component that copy and paste themselves into different genomic locations (transposon) by converting RNA back into DNA through the process reverse transcription using an RNA transposition intermediate.[1]
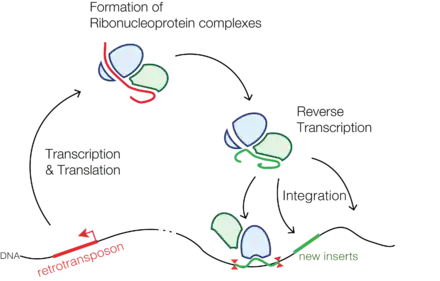
Through reverse transcription, retrotransposons amplify themselves quickly to become abundant in eukaryotic genomes such as maize (49–78%)[2] and humans (42%).[3] They are only present in eukaryotes but share features with retroviruses such as HIV, for example, discontinuous reverse transcriptase-mediated extrachromosomal recombination.[4][5]
There are two main types of retrotransposons, long terminal repeats (LTRs) and non-long terminal repeats (non-LTRs). Retrotransposons are classified based on sequence and method of transposition.[6] Most retrotransposons in the maize genome are LTR, whereas in humans they are mostly non-LTR. Retrotransposons (mostly of the LTR type) can be passed onto the next generation of a host species through the germline.
The other type of transposon is the DNA transposon. DNA transposons insert themselves into different genomic locations without copying themselves that can cause harmful mutations (see horizontal gene transfer). Hence retrotransposons can be thought of as replicative, whereas DNA transposons are non-replicative. Due to their replicative nature, retrotransposons can increase eukaryotic genome size quickly and survive in eukaryotic genomes permanently. It is thought that staying in eukaryotic genomes for such long periods gave rise to special insertion methods that do not affect eukaryotic gene function drastically.[7]
Replicative transposition
Using Tn3 plasmid as a model
LTR retrotransposons
Long strands of repetitive DNA can be found at each end of a LTR retrotransposon. These are termed long terminal repeats (LTRs) that are each a few hundred base pairs long, hence retrotransposons with LTRs have the name long terminal repeat (LTR) retrotransposon. LTR retrotransposons are over 5 kilobases long. Between the long terminal repeats there are genes that can be transcribed equivalent to retrovirus genes gag and pol. These genes overlap so they encode a protease that processes the resulting transcript into functional gene products. Gag gene products associate with other retrotransposon transcripts to form virus-like particles. Pol gene products include enzymes reverse transcriptase, integrase and ribonuclease H domains. Reverse transcriptase carries out reverse transcription of retrotransposon DNA. Integrase 'integrates' retrotransposon DNA into eukaryotic genome DNA. Ribonuclease cleaves phosphodiester bonds between RNA nucleotides.
LTR retrotransposons encode transcripts with tRNA binding sites so that they can undergo reverse transcription. The tRNA-bound RNA transcript binds to a genomic RNA sequence. Template strand of retrotransposon DNA can hence be synthesised. Ribonuclease H domains degrade eukaryotic genomic RNA to give adenine- and guanine-rich DNA sequences that flag where the complementary noncoding strand has to be synthesised. Integrase then 'integrates' the retrotransposon into eukaryotic DNA using the hydroxyl group at the start of retrotransposon DNA. This results in a retrotransposon flagged by long terminal repeats at its ends. Because the retrotransposon contains eukaryotic genome information it can insert copies of itself into other genomic locations within a eukaryotic cell.
Endogenous retrovirus
An endogenous retrovirus is a retrovirus without virus pathogenic effects that has been integrated into the host genome by inserting their inheritable genetic information into cells that can be passed onto the next generation like a retrotransposon.[8] Because of this, they share features with retroviruses and retrotransposons. When the retroviral DNA is integrated into the host genome they evolve into endogenous retroviruses that influence eukaryotic genomes. So many endogenous retroviruses have inserted themselves into eukaryotic genomes that they allow insight into biology between viral-host interactions and the role of retrotransposons in evolution and disease. Many retrotransposons share features with endogenous retroviruses, the property of recognising and fusing with the host genome. However, there is a key difference between retroviruses and retrotransposons, which is indicated by the env gene. Although similar to the gene carrying out the same function in retroviruses, the env gene is used to determine whether the gene is retroviral or retrotransposon. If the gene is retroviral it can evolve from a retrotransposon into a retrovirus. They differ by the order of sequences in pol genes. Env genes are found in LTR retrotransposon types Ty1-copia (Pseudoviridae), Ty3-gypsy (Metaviridae) and BEL/Pao.[9][8] They encode glycoproteins on the retrovirus envelope needed for entry into the host cell. Retroviruses can move between cells whereas LTR retrotransposons can only move themselves into the genome of the same cell.[10] Many vertebrate genes were formed from retroviruses and LTR retrotransposons. One endogenous retrovirus or LTR retrotransposon has the same function and genomic locations in different species, suggesting their role in evolution.[11]
Non-LTR retrotransposons
Like LTR retrotransposons, non-LTR retrotransposons contain genes for reverse transcriptase, RNA-binding protein, nuclease, and sometimes ribonuclease H domain[12] but they lack the long terminal repeats. RNA-binding proteins bind the RNA-transposition intermediate and nucleases are enzymes that break phosphodiester bonds between nucleotides in nucleic acids. Instead of LTRs, non-LTR retrotransposons have short repeats that can have an inverted order of bases next to each other aside from direct repeats found in LTR retrotransposons that is just one sequence of bases repeating itself.
Although they are retrotransposons, they cannot carry out reverse transcription using an RNA transposition intermediate in the same way as LTR retrotransposons. Those two key components of the retrotransposon are still necessary but the way they are incorporated into the chemical reactions is different. This is because unlike LTR retrotransposons, non-LTR retrotransposons do not contain sequences that bind tRNA.
They mostly fall into two types – LINEs and SINEs. SVA elements are the exception between the two as they share similarities with both LINEs and SINEs, containing Alu elements and different numbers of the same repeat. SVAs are shorter than LINEs but longer than SINEs.
While historically viewed as "junk DNA", research suggests in some cases, both LINEs and SINEs were incorporated into novel genes to form new functions.[13]
LINEs
When a LINE is transcribed, the transcript contains an RNA polymerase II promoter that ensures LINEs can be copied into whichever location it inserts itself into. RNA polymerase II is the enzyme that transcribes genes into mRNA transcripts. The ends of LINE transcripts are rich in multiple adenines,[14] the bases that are added at the end of transcription so that LINE transcripts would not be degraded. This transcript is the RNA transposition intermediate.
The RNA transposition intermediate moves from the nucleus into the cytoplasm for translation. This gives the two coding regions of a LINE that in turn binds back to the RNA it is transcribed from. The LINE RNA then moves back into the nucleus to insert into the eukaryotic genome.
LINEs insert themselves into regions of the eukaryotic genome that are rich in bases AT. At AT regions LINE uses its nuclease to cut one strand of the eukaryotic double-stranded DNA. The adenine-rich sequence in LINE transcript base pairs with the cut strand to flag where the LINE will be inserted with hydroxyl groups. Reverse transcriptase recognises these hydroxyl groups to synthesise LINE retrotransposon where the DNA is cut. Like with LTR retrotransposons, this new inserted LINE contains eukaryotic genome information so it can be copied and pasted into other genomic regions easily. The information sequences are longer and more variable than those in LTR retrotransposons.
Most LINE copies have variable length at the start because reverse transcription usually stops before DNA synthesis is complete. In some cases this causes RNA polymerase II promoter to be lost so LINEs cannot transpose further.[15]
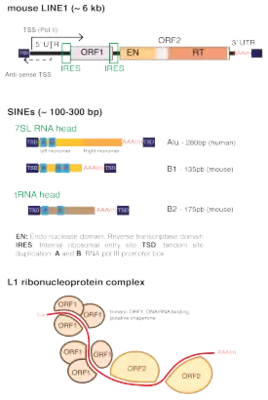
Human L1
LINE-1 (L1) retrotransposons make up a significant portion of the human genome, with an estimated 500,000 copies per genome. Genes encoding for human LINE1 usually have their transcription inhibited by methyl groups binding to its DNA carried out by PIWI proteins and enzymes DNA methyltransferases. L1 retrotransposition can disrupt the nature of genes transcribed by pasting themselves inside or near genes which could in turn lead to human disease. LINE1s can only retrotranspose in some cases to form different chromosome structures contributing to differences in genetics between individuals.[17] There is an estimate of 80–100 active L1s in the reference genome of the Human Genome Project, and an even smaller number of L1s within those active L1s retrotranspose often. L1 insertions have been associated with tumorigenesis by activating cancer-related genes oncogenes and tumor suppressors.
Each human LINE1 contains two regions from which gene products can be encoded. The first coding region contains a leucine zipper protein involved in protein-protein interactions and a protein that binds to the terminus of nucleic acids. The second coding region has a purine/pyrimidine nuclease, reverse transcriptase and protein rich in amino acids cysteines and histidines. The end of the human LINE1, as with other retrotransposons is adenine-rich.[18][19][20]
SINEs
SINEs are much shorter (300bp) than LINEs.[21] They share similarity with genes transcribed by RNA polymerase II, the enzyme that transcribes genes into mRNA transcripts, and the initiation sequence of RNA polymerase III, the enzyme that transcribes genes into ribosomal RNA, tRNA and other small RNA molecules.[22] SINEs such as mammalian MIR elements have tRNA gene at the start and adenine-rich at the end like in LINEs.
SINEs do not encode a functional reverse transcriptase protein and rely on other mobile transposons, especially LINEs.[23] SINEs exploit LINE transposition components despite LINE-binding proteins prefer binding to LINE RNA. SINEs cannot transpose by themselves because they cannot encode SINE transcripts. They usually consist of parts derived from tRNA and LINEs. The tRNA portion contains an RNA polymerase III promoter which the same kind of enzyme as RNA polymerase II. This makes sure the LINE copies would be transcribed into RNA for further transposition. The LINE component remains so LINE-binding proteins can recognise the LINE part of the SINE.
Alu elements
Alus are the most common SINE in primates. They are approximately 350 base pairs long, do not encode proteins and can be recognized by the restriction enzyme AluI (hence the name). Their distribution may be important in some genetic diseases and cancers. Copy and pasting Alu RNA requires the Alu's adenine-rich end and the rest of the sequence bound to a signal. The signal-bound Alu can then associate with ribosomes. LINE RNA associates on the same ribosomes as the Alu. Binding to the same ribosome allows Alus of SINEs to interact with LINE. This simultaneous translation of Alu element and LINE allows SINE copy and pasting.
SVA elements
SVA elements are present at lower levels than SINES and LINEs in humans. The starts of SVA and Alu elements are similar, followed by repeats and an end similar to endogenous retrovirus. LINEs bind to sites flanking SVA elements to transpose them. SVA are one of the youngest transposons in great apes genome and among the most active and polymorphic in the human population.
A recent study developed a network method that reveals SVA retroelement (RE) proliferation dynamics in hominid genomes.[24] The method enable to track the course of SVA proliferation, identify yet unknown active communities, and detect tentative "master REs" that play key roles in SVA propagation. Thus, providing support for the fundamental "master gene" model of RE proliferation.
Role in human disease
Retrotransposons ensure they are not lost by chance by occurring only in cell genetics that can be passed on from one generation to the next from parent gametes. However, LINEs can transpose into the human embryo cells that eventually develop into the nervous system, raising the question whether this LINE retrotransposition affects brain function. LINE retrotransposition is also a feature of several cancers, but it is unclear whether retrotransposition itself causes cancer instead of just a symptom. Uncontrolled retrotransposition is bad for both the host organism and retrotransposons themselves so they have to be regulated. Retrotransposons are regulated by RNA interference. RNA interference is carried out by a bunch of short non-coding RNAs. The short non-coding RNA interacts with protein Argonaute to degrade retrotransposon transcripts and change their DNA histone structure to reduce their transcription.
Role in evolution
LTR retrotransposons came about later than non-LTR retrotransposons, possibly from an ancestral non-LTR retrotransposon acquiring an integrase from a DNA transposon. Retroviruses gained additional properties to their virus envelopes by taking the relevant genes from other viruses using the power of LTR retrotransposon.
Due to their retrotransposition mechanism, retrotransposons amplify in number quickly, composing 40% of the human genome. The insertion rates for LINE1, Alu and SVA elements are 1/200 – 1/20, 1/20 and 1/900 respectively. The LINE1 insertion rates have varied a lot over the past 35 million years, so they indicate points in genome evolution.
Notably a large number of 100 kilobases in the maize genome show variety due to the presence or absence of retrotransposons. However since maize is unusually genetically compared to other plants it cannot be used to predict retrotransposition in other plants.
Mutations caused by retrotransposons include:
- Gene inactivation
- Changing gene regulation
- Changing gene products
- Acting as DNA repair sites
See also
- Copy-number variation
- Genomic organization
- Insertion sequences
- Interspersed repeat
- Paleogenetics
- Paleovirology
- RetrOryza
- Retrotransposon markers, a powerful method of reconstructing phylogenies.
- Tn3 transposon
- Transposon
References
- Dombroski BA, Feng Q, Mathias SL, Sassaman DM, Scott AF, Kazazian HH, Boeke JD (July 1994). "An in vivo assay for the reverse transcriptase of human retrotransposon L1 in Saccharomyces cerevisiae". Molecular and Cellular Biology. 14 (7): 4485–92. doi:10.1128/mcb.14.7.4485. PMC 358820. PMID 7516468.
- SanMiguel P, Bennetzen JL (1998). "Evidence that a recent increase in maize genome size was caused by the massive amplification of intergene retrotranposons" (PDF). Annals of Botany. 82 (Suppl A): 37–44. doi:10.1006/anbo.1998.0746.
- Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, et al. (February 2001). "Initial sequencing and analysis of the human genome". Nature. 409 (6822): 860–921. Bibcode:2001Natur.409..860L. doi:10.1038/35057062. PMID 11237011.
- Sanchez, Diego; Gaubert, Hervé; Drost, Hajk-Georg; Zabet, Nicolae Radu; Paszkowski, Jerzy (2017-11-03). "High-frequency recombination between members of an LTR retrotransposon family during transposition bursts". Nature Communications. 8: 11283. doi:10.1038/s41467-017-01374-x. PMC 5668417. PMID 29097664.
- Drost, Hajk-Georg; Sanchez, Diego (2019-12-01). "Becoming a selfish clan: recombination associated to reverse-transcription in LTR retrotransposons". Genome Biology and Evolution. 11 (12): 3382–3392. doi:10.1093/gbe/evz255. PMC 6894440. PMID 31755923.
- Xiong Y, Eickbush TH (October 1990). "Origin and evolution of retroelements based upon their reverse transcriptase sequences". The EMBO Journal. 9 (10): 3353–62. doi:10.1002/j.1460-2075.1990.tb07536.x. PMC 552073. PMID 1698615.
- Finnegan DJ (June 2012). "Retrotransposons". Cell Current Biology. 22 (11): R432–R437. doi:10.1016/j.cub.2012.04.025. PMID 22677280.
- Wicker T, Sabot F, Hua-Van A, Bennetzen JL, Capy P, Chalhoub B, Flavell A, Leroy P, Morgante M, Panaud O, Paux E, SanMiguel P, Schulman AH (December 2007). "A unified classification system for eukaryotic transposable elements". Nature Reviews. Genetics. 8 (12): 973–82. doi:10.1038/nrg2165. PMID 17984973. S2CID 32132898.
- Copeland CS, Mann VH, Morales ME, Kalinna BH, Brindley PJ (February 2005). "The Sinbad retrotransposon from the genome of the human blood fluke, Schistosoma mansoni, and the distribution of related Pao-like elements". BMC Evolutionary Biology. 5 (1): 20. doi:10.1186/1471-2148-5-20. PMC 554778. PMID 15725362.
- Havecker ER, Gao X, Voytas DF (18 May 2004). "The diversity of LTR retrotransposons". BMC Genome Biology. 5 (225): 225. doi:10.1186/gb-2004-5-6-225. PMC 463057. PMID 15186483.
- Naville M, Warren IA, Haftek-Terreau Z, Chalopin D, Brunet F, Levin P, Galiana D, Volff JN (17 February 2016). "Not so bad after all: retroviruses and long terminal repeat retrotransposons as a source of new genes in vertebrates". Clinical Microbiology and Infection. 22 (4): 312–323. doi:10.1016/j.cmi.2016.02.001. PMID 26899828.
- Yadav VP, Mandal PK, Rao DN, Bhattacharya S (December 2009). "Characterization of the restriction enzyme-like endonuclease encoded by the Entamoeba histolytica non-long terminal repeat retrotransposon EhLINE1". The FEBS Journal. 276 (23): 7070–82. doi:10.1111/j.1742-4658.2009.07419.x. PMID 19878305.
- Santangelo AM, de Souza FS, Franchini LF, Bumaschny VF, Low MJ, Rubinstein M (October 2007). "Ancient exaptation of a CORE-SINE retroposon into a highly conserved mammalian neuronal enhancer of the proopiomelanocortin gene". PLOS Genetics. 3 (10): 1813–26. doi:10.1371/journal.pgen.0030166. PMC 2000970. PMID 17922573.
- Liang KH, Yeh CT (May 2013). "A gene expression restriction network mediated by sense and antisense Alu sequences located on protein-coding messenger RNAs". BMC Genomics. 14: 325. doi:10.1186/1471-2164-14-325. PMC 3655826. PMID 23663499.
- Singer MF (March 1982). "SINEs and LINEs: highly repeated short and long interspersed sequences in mammalian genomes". Cell. 28 (3): 433–4. doi:10.1016/0092-8674(82)90194-5. PMID 6280868. S2CID 22129236.
- Walter M (2015). "Transposon regulation upon dynamic loss of DNA methylation (PDF Download Available)". ResearchGate. doi:10.13140/rg.2.2.18747.21286.
- Chueh AC, Northrop EL, Brettingham-Moore KH, Choo KH, Wong LH (January 2009). Bickmore WA (ed.). "LINE retrotransposon RNA is an essential structural and functional epigenetic component of a core neocentromeric chromatin". PLOS Genetics. 5 (1): e1000354. doi:10.1371/journal.pgen.1000354. PMC 2625447. PMID 19180186.
- Doucet AJ, Hulme AE, Sahinovic E, Kulpa DA, Moldovan JB, Kopera HC, Athanikar JN, Hasnaoui M, Bucheton A, Moran JV, Gilbert N (October 2010). "Characterization of LINE-1 ribonucleoprotein particles". PLOS Genetics. 6 (10): e1001150. doi:10.1371/journal.pgen.1001150. PMC 2951350. PMID 20949108.
- Denli AM, Narvaiza I, Kerman BE, Pena M, Benner C, Marchetto MC, Diedrich JK, Aslanian A, Ma J, Moresco JJ, Moore L, Hunter T, Saghatelian A, Gage FH (October 2015). "Primate-specific ORF0 contributes to retrotransposon-mediated diversity". Cell. 163 (3): 583–93. doi:10.1016/j.cell.2015.09.025. PMID 26496605. S2CID 10525450.
- Ohshima K, Okada N (2005). "SINEs and LINEs: symbionts of eukaryotic genomes with a common tail". Cytogenetic and Genome Research. 110 (1–4): 475–90. doi:10.1159/000084981. PMID 16093701. S2CID 42841487.
- Stansfield WD, King RC (1997). A dictionary of genetics (5th ed.). Oxford [Oxfordshire]: Oxford University Press. ISBN 978-0-19-509441-1.
- Kramerov DA, Vassetzky NS (2005). "Short retroposons in eukaryotic genomes". International Review of Cytology. 247: 165–221. doi:10.1016/s0074-7696(05)47004-7. PMID 16344113.
- Dewannieux M, Esnault C, Heidmann T (September 2003). "LINE-mediated retrotransposition of marked Alu sequences". Nature Genetics. 35 (1): 41–8. doi:10.1038/ng1223. PMID 12897783. S2CID 32151696.
- Levy OR, Knisbacher BA, Levanon EY, Havlin SH (2017). "Integrating networks and comparative genomics reveals retroelement proliferation dynamics in hominid genomes". Science Advances. 3 (7): e1701256. Bibcode:2017SciA....3E1256L. doi:10.1126/sciadv.1701256. PMC 5640379. PMID 29043294.