Catalase
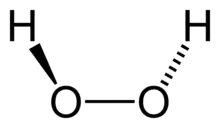
Catalase is a common enzyme found in nearly all living organisms exposed to oxygen (such as bacteria, plants, and animals) which catalyzes the decomposition of hydrogen peroxide to water and oxygen.[1] It is a very important enzyme in protecting the cell from oxidative damage by reactive oxygen species (ROS). Likewise, catalase has one of the highest turnover numbers of all enzymes; one catalase molecule can convert millions of hydrogen peroxide molecules to water and oxygen each second.[2]
| Catalase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| Symbol | Catalase | ||||||||
| Pfam | PF00199 | ||||||||
| InterPro | IPR011614 | ||||||||
| PROSITE | PDOC00395 | ||||||||
| SCOP2 | 7cat / SCOPe / SUPFAM | ||||||||
| OPM superfamily | 370 | ||||||||
| OPM protein | 3e4w | ||||||||
| CDD | cd00328 | ||||||||
| |||||||||
| catalase | |||||||
|---|---|---|---|---|---|---|---|
 | |||||||
| Identifiers | |||||||
| Aliases | IPR024708Catalase active siteprotein family | ||||||
| External IDs | GeneCards: | ||||||
| Orthologs | |||||||
| Species | Human | Mouse | |||||
| Entrez |
|
| |||||
| Ensembl |
|
| |||||
| UniProt |
|
| |||||
| RefSeq (mRNA) |
|
| |||||
| RefSeq (protein) |
|
| |||||
| Location (UCSC) | n/a | n/a | |||||
| PubMed search | n/a | n/a | |||||
| Wikidata | |||||||
| |||||||
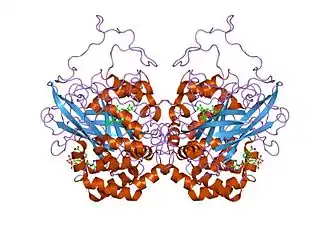
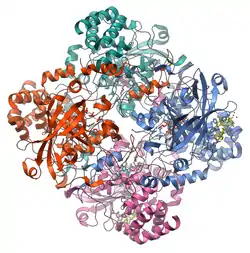
Catalase is a tetramer of four polypeptide chains, each over 500 amino acids long.[3] It contains four iron-containing heme groups that allow the enzyme to react with the hydrogen peroxide. The optimum pH for human catalase is approximately 7,[4] and has a fairly broad maximum: the rate of reaction does not change appreciably between pH 6.8 and 7.5.[5] The pH optimum for other catalases varies between 4 and 11 depending on the species.[6] The optimum temperature also varies by species.[7]
Structure
Human catalase forms a tetramer composed of four subunits, each of which can be conceptually divided into four domains.[8] The extensive core of each subunit is generated by an eight-stranded antiparallel b-barrel (b1-8), with nearest neighbor connectivity capped by b-barrel loops on one side and a9 loops on the other.[8] A helical domain at one face of the b-barrel is composed of four C-terminal helices (a16, a17, a18, and a19) and four helices derived from residues between b4 and b5 (a4, a5, a6, and a7).[8] Alternative splicing may result in different protein variants.
History
Catalase was first noticed in 1818 when Louis Jacques Thénard, who discovered H2O2 (hydrogen peroxide), suggested its breakdown is caused by an unknown substance. In 1900, Oscar Loew was the first to give it the name catalase, and found it in many plants and animals.[9] In 1937 catalase from beef liver was crystallised by James B. Sumner and Alexander Dounce[10] and the molecular weight was found in 1938.[11]
The amino acid sequence of bovine catalase was determined in 1969,[12] and the three-dimensional structure in 1981.[13]
Function
Reaction
- 2 H2O2 → 2 H2O + O2
The presence of catalase in a microbial or tissue sample can be demonstrated by adding hydrogen peroxide and observing the reaction. The production of oxygen can be seen by the formation of bubbles. This easy test, which can be seen with the naked eye, without the aid of instruments, is possible because catalase has a very high specific activity, which produces a detectable response, as well as the fact that one of the products is a gas.
Molecular mechanism
While the complete mechanism of catalase is not currently known,[14] the reaction is believed to occur in two stages:
- H2O2 + Fe(III)-E → H2O + O=Fe(IV)-E(.+)
- H2O2 + O=Fe(IV)-E(.+) → H2O + Fe(III)-E + O2[14]
Here Fe()-E represents the iron center of the heme group attached to the enzyme. Fe(IV)-E(.+) is a mesomeric form of Fe(V)-E, meaning the iron is not completely oxidized to +V, but receives some stabilising electron density from the heme ligand, which is then shown as a radical cation (.+).
As hydrogen peroxide enters the active site, it interacts with the amino acids Asn148 (asparagine at position 148) and His75, causing a proton (hydrogen ion) to transfer between the oxygen atoms. The free oxygen atom coordinates, freeing the newly formed water molecule and Fe(IV)=O. Fe(IV)=O reacts with a second hydrogen peroxide molecule to reform Fe(III)-E and produce water and oxygen.[14] The reactivity of the iron center may be improved by the presence of the phenolate ligand of Tyr358 in the fifth coordination position, which can assist in the oxidation of the Fe(III) to Fe(IV). The efficiency of the reaction may also be improved by the interactions of His75 and Asn148 with reaction intermediates.[14] The decomposition of hydrogen peroxide by catalase proceeds according to first-order kinetics, the rate being proportional to the hydrogen peroxide concentration.[15]
Catalase can also catalyze the oxidation, by hydrogen peroxide, of various metabolites and toxins, including formaldehyde, formic acid, phenols, acetaldehyde and alcohols. It does so according to the following reaction:
- H2O2 + H2R → 2H2O + R
The exact mechanism of this reaction is not known.
Any heavy metal ion (such as copper cations in copper(II) sulfate) can act as a noncompetitive inhibitor of catalase. Furthermore, the poison cyanide is a noncompetitive inhibitor[16] of catalase at high concentrations of hydrogen peroxide.[17] Arsenate acts as an activator.[18] Three-dimensional protein structures of the peroxidated catalase intermediates are available at the Protein Data Bank.
Cellular role
Hydrogen peroxide is a harmful byproduct of many normal metabolic processes; to prevent damage to cells and tissues, it must be quickly converted into other, less dangerous substances. To this end, catalase is frequently used by cells to rapidly catalyze the decomposition of hydrogen peroxide into less-reactive gaseous oxygen and water molecules.[19]
Mice genetically engineered to lack catalase are initially phenotypically normal.,[20] however, catalase deficiency in mice may increase the likelihood of developing obesity, fatty liver,[21] and type 2 diabetes.[22] Some humans have very low levels of catalase (acatalasia), yet show few ill effects.
The increased oxidative stress that occurs with aging in mice is alleviated by over-expression of catalase.[23] Over-expressing mice do not exhibit the age-associated loss of spermatozoa, testicular germ and Sertoli cells seen in wild-type mice. Oxidative stress in wild-type mice ordinarily induces oxidative DNA damage (measured as 8-oxodG) in sperm with aging, but these damages are significantly reduced in aged catalase over-expressing mice.[23] Furthermore, these over-expressing mice show no decrease in age-dependent number of pups per litter. Overexpression of catalase targeted to mitochondria extends the lifespan of mice.[24]
Catalase is usually located in a cellular organelle called the peroxisome.[25] Peroxisomes in plant cells are involved in photorespiration (the use of oxygen and production of carbon dioxide) and symbiotic nitrogen fixation (the breaking apart of diatomic nitrogen (N2) to reactive nitrogen atoms). Hydrogen peroxide is used as a potent antimicrobial agent when cells are infected with a pathogen. Catalase-positive pathogens, such as Mycobacterium tuberculosis, Legionella pneumophila, and Campylobacter jejuni, make catalase to deactivate the peroxide radicals, thus allowing them to survive unharmed within the host.[26]
Like alcohol dehydrogenase, catalase converts ethanol to acetaldehyde, but it is unlikely that this reaction is physiologically significant.[27]
Distribution among organisms
The large majority of known organisms use catalase in every organ, with particularly high concentrations occurring in the liver in mammals.[28] Catalase is found primarily in peroxisomes and the cytosol of erythrocytes (and sometimes in mitochondria[29])
Almost all aerobic microorganisms use catalase. It is also present in some anaerobic microorganisms, such as Methanosarcina barkeri.[30] Catalase is also universal among plants and occurs in most fungi.[31]
One unique use of catalase occurs in the bombardier beetle. This beetle has two sets of liquids that are stored separately in two paired glands. The larger of the pair, the storage chamber or reservoir, contains hydroquinones and hydrogen peroxide, while the smaller, the reaction chamber, contains catalases and peroxidases. To activate the noxious spray, the beetle mixes the contents of the two compartments, causing oxygen to be liberated from hydrogen peroxide. The oxygen oxidizes the hydroquinones and also acts as the propellant.[32] The oxidation reaction is very exothermic (ΔH = −202.8 kJ/mol) and rapidly heats the mixture to the boiling point.[33]
Long-lived queens of the termite Reticulitermes speratus have significantly lower oxidative damage to their DNA than non-reproductive individuals (workers and soldiers).[34] Queens have more than two times higher catalase activity and seven times higher expression levels of the catalase gene RsCAT1 than workers.[34] It appears that the efficient antioxidant capability of termite queens can partly explain how they attain longer life.
Catalase enzymes from various species have vastly differing optimum temperatures. Poikilothermic animals typically have catalases with optimum temperatures in the range of 15-25 °C, while mammalian or avian catalases might have optimum temperatures above 35 °C,[35][36] and catalases from plants vary depending on their growth habit.[35] In contrast, catalase isolated from the hyperthermophile archaeon Pyrobaculum calidifontis has a temperature optimum of 90 °C.[37]
Clinical significance and application
Catalase is used in the food industry for removing hydrogen peroxide from milk prior to cheese production.[38] Another use is in food wrappers where it prevents food from oxidizing.[39] Catalase is also used in the textile industry, removing hydrogen peroxide from fabrics to make sure the material is peroxide-free.[40]
A minor use is in contact lens hygiene – a few lens-cleaning products disinfect the lens using a hydrogen peroxide solution; a solution containing catalase is then used to decompose the hydrogen peroxide before the lens is used again.[41]
Bacterial identification (catalase test)

The catalase test is one of the three main tests used by microbiologists to identify species of bacteria. If the bacteria possess catalase (i.e., are catalase-positive), when a small amount of bacterial isolate is added to hydrogen peroxide, bubbles of oxygen are observed. The catalase test is done by placing a drop of hydrogen peroxide on a microscope slide. An applicator stick is touched to the colony, and the tip is then smeared onto the hydrogen peroxide drop.
- If the mixture produces bubbles or froth, the organism is said to be 'catalase-positive'. Staphylococci[42] and Micrococci[43] are catalase-positive. Other catalase-positive organisms include Listeria, Corynebacterium diphtheriae, Burkholderia cepacia, Nocardia, the family Enterobacteriaceae (Citrobacter, E. coli, Enterobacter, Klebsiella, Shigella, Yersinia, Proteus, Salmonella, Serratia), Pseudomonas, Mycobacterium tuberculosis, Aspergillus, Cryptococcus, and Rhodococcus equi.
- If not, the organism is 'catalase-negative'. Streptococcus[44] and Enterococcus spp. are catalase-negative.
While the catalase test alone cannot identify a particular organism, it can aid identification when combined with other tests such as antibiotic resistance. The presence of catalase in bacterial cells depends on both the growth condition and the medium used to grow the cells.
Capillary tubes may also be used. A small sample of bacteria is collected on the end of the capillary tube, without blocking the tube, to avoid false negative results. The opposite end is then dipped into hydrogen peroxide, which is drawn into the tube through capillary action, and turned upside down, so that the bacterial sample points downwards. The hand holding the tube is then tapped on the bench, moving the hydrogen peroxide down until it touches the bacteria. If bubbles form on contact, this indicates a positive catalase result. This test can detect catalase-positive bacteria at concentrations above about 105 cells/mL,[45] and is simple to use.
Bacterial virulence
Neutrophils and other phagocytes use peroxide to kill bacteria. The enzyme NADPH oxidase generates superoxide within the phagosome, which is converted via hydrogen peroxide to other oxidising substances like hypochlorous acid which kill phagocytosed pathogens.[46] In individuals with chronic granulomatous disease (CGD) there is a defect in producing peroxide via mutations in phagocyte oxidases such as myeloperoxidase.[47] Normal cellular metabolism will still produce a small amount of peroxide and this peroxide can be used to produce hypochlorous acid to eradicate the bacterial infection. However, if individuals with CGD are infected with catalase-positive bacteria, the bacterial catalase can destroy the excess peroxide before it can be used to produce other oxidising substances. In these individuals the pathogen survives and becomes a chronic infection. This chronic infection is typically surrounded by macrophages in an attempt to isolate the infection. This wall of macrophages surrounding a pathogen is called a granuloma. Many bacteria are catalase positive, but some are better catalase-producers than others. The mnemonic "cats Need PLACESS to Belch their Hairballs" can be used to memorise the catalase-positive bacteria (and Candida and Aspergillus, which are fungi): nocardia, pseudomonas, listeria, aspergillus, candida, E. coli, staphylococcus, serratia, B. cepacia and H. pylori.[48]
Acatalasia
Acatalasia is a condition caused by homozygous mutations in CAT, resulting in a lack of catalase. Symptoms are mild and include oral ulcers. A heterozygous CAT mutation results in lower, but still present catalase.[49]
Gray hair
Low levels of catalase may play a role in the graying process of human hair. Hydrogen peroxide is naturally produced by the body and broken down by catalase. If catalase levels decline, hydrogen peroxide cannot be broken down so well. The hydrogen peroxide interferes with the production of melanin, the pigment that gives hair its color.[50][51]
Interactions
Catalase has been shown to interact with the ABL2[52] and Abl genes.[52] Infection with the murine leukemia virus causes catalase activity to decline in the lungs, heart and kidneys of mice. Conversely, dietary fish oil increased catalase activity in the heart, and kidneys of mice.[53]
References
- Chelikani P, Fita I, Loewen PC (January 2004). "Diversity of structures and properties among catalases". Cellular and Molecular Life Sciences. 61 (2): 192–208. doi:10.1007/s00018-003-3206-5. hdl:10261/111097. PMID 14745498. S2CID 4411482.
- Goodsell DS (2004-09-01). "Catalase". Molecule of the Month. RCSB Protein Data Bank. Retrieved 2016-08-23.
- Boon EM, Downs A, Marcey D. "Catalase: H2O2: H2O2 Oxidoreductase". Catalase Structural Tutorial Text. Retrieved 2007-02-11.
- Maehly AC, Chance B (1954). "The assay of catalases and peroxidases". Methods of Biochemical Analysis. Methods of Biochemical Analysis. 1. pp. 357–424. doi:10.1002/9780470110171.ch14. ISBN 978-0-470-11017-1. PMID 13193536.
- Aebi H (1984). Catalase in vitro. Methods in Enzymology. 105. pp. 121–6. doi:10.1016/S0076-6879(84)05016-3. ISBN 978-0-12-182005-3. PMID 6727660.
- "EC 1.11.1.6 - catalase". BRENDA: The Comprehensive Enzyme Information System. Department of Bioinformatics and Biochemistry, Technische Universität Braunschweig. Retrieved 2009-05-26.
- Toner K, Sojka G, Ellis R. "A Quantitative Enzyme Study; CATALASE". bucknell.edu. Archived from the original on 2000-06-12. Retrieved 2007-02-11.
- Putnam CD, Arvai AS, Bourne Y, Tainer JA (February 2000). "Active and inhibited human catalase structures: ligand and NADPH binding and catalytic mechanism". Journal of Molecular Biology. 296 (1): 295–309. doi:10.1006/jmbi.1999.3458. PMID 10656833.
- Loew O (May 1900). "A New Enzyme of General Occurrence in Organisms". Science. 11 (279): 701–2. Bibcode:1900Sci....11..701L. doi:10.1126/science.11.279.701. JSTOR 1625707. PMID 17751716.
- Sumner JB, Dounce AL (April 1937). "Crystalline Catalase". Science. 85 (2206): 366–7. Bibcode:1937Sci....85..366S. doi:10.1126/science.85.2206.366. PMID 17776781.
- Sumner JB, Gralén N (March 1938). "The Molecular Weight Of Crystalline Catalase". Science. 87 (2256): 284. Bibcode:1938Sci....87..284S. doi:10.1126/science.87.2256.284. PMID 17831682.
- Schroeder WA, Shelton JR, Shelton JB, Robberson B, Apell G (May 1969). "The amino acid sequence of bovine liver catalase: a preliminary report". Archives of Biochemistry and Biophysics. 131 (2): 653–5. doi:10.1016/0003-9861(69)90441-X. PMID 4892021.
- Murthy MR, Reid TJ, Sicignano A, Tanaka N, Rossmann MG (October 1981). "Structure of beef liver catalase". Journal of Molecular Biology. 152 (2): 465–99. doi:10.1016/0022-2836(81)90254-0. PMID 7328661.
- Boon EM, Downs A, Marcey D. "Proposed Mechanism of Catalase". Catalase: H2O2: H2O2 Oxidoreductase: Catalase Structural Tutorial. Retrieved 2007-02-11.
- Aebi H (1984). "Catalase in vitro". Methods in Enzymology. 105: 121–6. doi:10.1016/S0076-6879(84)05016-3. PMID 6727660.
- Nonstationary Inhibition of Enzyme Action. The Cyanide Inhibition of Catalase
- Ogura Y, Yamazaki I (August 1983). "Steady-state kinetics of the catalase reaction in the presence of cyanide". Journal of Biochemistry. 94 (2): 403–8. doi:10.1093/oxfordjournals.jbchem.a134369. PMID 6630165.
- Kertulis-Tartar GM, Rathinasabapathi B, Ma LQ (October 2009). "Characterization of glutathione reductase and catalase in the fronds of two Pteris ferns upon arsenic exposure". Plant Physiology and Biochemistry. 47 (10): 960–5. doi:10.1016/j.plaphy.2009.05.009. PMID 19574057.
- Gaetani GF, Ferraris AM, Rolfo M, Mangerini R, Arena S, Kirkman HN (February 1996). "Predominant role of catalase in the disposal of hydrogen peroxide within human erythrocytes". Blood. 87 (4): 1595–9. doi:10.1182/blood.V87.4.1595.bloodjournal8741595. PMID 8608252.
- Ho YS, Xiong Y, Ma W, Spector A, Ho DS (July 2004). "Mice lacking catalase develop normally but show differential sensitivity to oxidant tissue injury". The Journal of Biological Chemistry. 279 (31): 32804–12. doi:10.1074/jbc.M404800200. PMID 15178682.
- Heit C, Marshall S, Singh S, Yu X, Charkoftaki G, Zhao H, Orlicky DJ, Fritz KS, Thompson DC, Vasiliou V (2017). "Catalase deletion promotes prediabetic phenotype in mice". Free Radical Biology & Medicine. 103: 48–56. doi:10.1016/j.freeradbiomed.2016.12.011. PMC 5513671. PMID 27939935.
- Góth L, Nagy T (2012). "Acatalasemia and diabetes mellitus". Archives of Biochemistry and Biophysics. 525 (2): 195–200. doi:10.1016/j.abb.2012.02.005. PMID 22365890.
- Selvaratnam J, Robaire B (November 2016). "Overexpression of catalase in mice reduces age-related oxidative stress and maintains sperm production". Exp. Gerontol. 84: 12–20. doi:10.1016/j.exger.2016.08.012. PMID 27575890. S2CID 2416413.
- Schriner SE, Linford NJ, Martin GM, Treuting P, Ogburn CE, Emond M, Coskun PE, Ladiges W, Wolf N, Van Remmen H, Wallace DC, Rabinovitch PS (June 2005). "Extension of murine life span by overexpression of catalase targeted to mitochondria". Science. 308 (5730): 1909–11. Bibcode:2005Sci...308.1909S. doi:10.1126/science.1106653. PMID 15879174. S2CID 38568666.
- Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). "Peroxisomes". Molecular Biology of the Cell (4th ed.). New York: Garland Science. ISBN 978-0-8153-3218-3.
- Srinivasa Rao PS, Yamada Y, Leung KY (September 2003). "A major catalase (KatB) that is required for resistance to H2O2 and phagocyte-mediated killing in Edwardsiella tarda". Microbiology. 149 (Pt 9): 2635–44. doi:10.1099/mic.0.26478-0. PMID 12949187.
- Lieber, Charles S. (January 1997). "Ethanol metabolism, cirrhosis and alcoholism". Clinica Chimica Acta. 257 (1): 59–84. doi:10.1016/S0009-8981(96)06434-0. PMID 9028626.
- Ilyukha VA (2001). "Superoxide Dismutase and Catalase in the Organs of Mammals of Different Ecogenesis". Journal of Evolutionary Biochemistry and Physiology. 37 (3): 241–245. doi:10.1023/A:1012663105999. S2CID 38916410.
- Bai J, Cederbaum AI (2001). "Mitochondrial catalase and oxidative injury". Biological Signals and Receptors. 10 (3–4): 3189–199. doi:10.1159/000046887. PMID 11351128. S2CID 33795198.
- Brioukhanov AL, Netrusov AI, Eggen RI (June 2006). "The catalase and superoxide dismutase genes are transcriptionally up-regulated upon oxidative stress in the strictly anaerobic archaeon Methanosarcina barkeri". Microbiology. 152 (Pt 6): 1671–7. doi:10.1099/mic.0.28542-0. PMID 16735730.
- Hansberg W, Salas-Lizana R, Domínguez L (September 2012). "Fungal catalases: function, phylogenetic origin and structure". Archives of Biochemistry and Biophysics. 525 (2): 170–80. doi:10.1016/j.abb.2012.05.014. PMID 22698962.
- Eisner T, Aneshansley DJ (August 1999). "Spray aiming in the bombardier beetle: photographic evidence". Proceedings of the National Academy of Sciences of the United States of America. 96 (17): 9705–9. Bibcode:1999PNAS...96.9705E. doi:10.1073/pnas.96.17.9705. PMC 22274. PMID 10449758.
- Beheshti N, McIntosh AC (2006). "A biomimetic study of the explosive discharge of the bombardier beetle" (PDF). Int. Journal of Design & Nature. 1 (1): 1–9. Archived from the original (PDF) on 2011-07-26.
- Tasaki E, Kobayashi K, Matsuura K, Iuchi Y (2017). "An Efficient Antioxidant System in a Long-Lived Termite Queen". PLOS ONE. 12 (1): e0167412. Bibcode:2017PLoSO..1267412T. doi:10.1371/journal.pone.0167412. PMC 5226355. PMID 28076409.
- Mitsuda, Hisateru (1956-07-31). "Studies on Catalase" (PDF). Bulletin of the Institute for Chemical Research, Kyoto University. 34 (4): 165–192. Retrieved 27 September 2017.
- Akkuş Çetinus Ş, Nursevin Öztop H (June 2003). "Immobilization of catalase into chemically crosslinked chitosan beads". Enzyme and Microbial Technology. 32 (7): 889–894. doi:10.1016/S0141-0229(03)00065-6.
- Amo T, Atomi H, Imanaka T (June 2002). "Unique presence of a manganese catalase in a hyperthermophilic archaeon, Pyrobaculum calidifontis VA1". Journal of Bacteriology. 184 (12): 3305–12. doi:10.1128/JB.184.12.3305-3312.2002. PMC 135111. PMID 12029047.
- "Catalase". Worthington Enzyme Manual. Worthington Biochemical Corporation. Retrieved 2009-03-01.
- Hengge A (1999-03-16). "Re: how is catalase used in industry?". General Biology. MadSci Network. Retrieved 2009-03-01.
- "textile industry". Case study 228. International Cleaner Production Information Clearinghouse. Archived from the original on 2008-11-04. Retrieved 2009-03-01.
- US patent 5521091, Cook JN, Worsley JL, "Compositions and method for destroying hydrogen peroxide on contact lens", issued 1996-05-28
- Rollins DM (2000-08-01). "Bacterial Pathogen List". BSCI 424 Pathogenic Microbiology. University of Maryland. Retrieved 2009-03-01.
- Johnson M. "Catalase Production". Biochemical Tests. Mesa Community College. Archived from the original on 2008-12-11. Retrieved 2009-03-01.
- Fox A. "Streptococcus pneumoniae and Staphylococci". University of South Carolina. Retrieved 2009-03-01.
- Martin, A. M. (2012-12-06). Fisheries Processing: Biotechnological applications. Springer Science & Business Media. ISBN 9781461553038.
- Winterbourn, Christine C.; Kettle, Anthony J.; Hampton, Mark B. (2016-06-02). "Reactive Oxygen Species and Neutrophil Function". Annual Review of Biochemistry. 85 (1): 765–792. doi:10.1146/annurev-biochem-060815-014442. ISSN 0066-4154. PMID 27050287.
- Murphy, Patrick (2012-12-06). The Neutrophil. Springer Science & Business Media. ISBN 9781468474183.
- Le, Tao; Bhushan, Vikas (2017-01-06). First aid for the USMLE step 1 2017 : a student-to-student guide. Le, Tao,, Bhushan, Vikas,, Sochat, Matthew,, Kallianos, Kimberly,, Chavda, Yash,, Zureick, Andrew H. (Andrew Harrison), 1991- (27th ed.). New York. ISBN 9781259837623. OCLC 986222844.
- "OMIM Entry - # 614097 - ACATALASEMIA". www.omim.org.
- "Why Hair Turns Gray Is No Longer A Gray Area: Our Hair Bleaches Itself As We Grow Older". Science News. ScienceDaily. 2009-02-24. Retrieved 2009-03-01.
- Wood JM, Decker H, Hartmann H, Chavan B, Rokos H, Spencer JD, Hasse S, Thornton MJ, Shalbaf M, Paus R, Schallreuter KU (July 2009). "Senile hair graying: H2O2-mediated oxidative stress affects human hair color by blunting methionine sulfoxide repair". FASEB Journal. 23 (7): 2065–75. arXiv:0706.4406. doi:10.1096/fj.08-125435. PMID 19237503.
- Cao C, Leng Y, Kufe D (August 2003). "Catalase activity is regulated by c-Abl and Arg in the oxidative stress response". The Journal of Biological Chemistry. 278 (32): 29667–75. doi:10.1074/jbc.M301292200. PMID 12777400.
- Xi S, Chen LH (2000). "Effects of dietary fish oil on tissue glutathione and antioxidant defense enzymes in mice with murine aids". Nutrition Research. 20 (9): 1287–99. doi:10.1016/S0271-5317(00)00214-1.
External links
- "GenAge entry for CAT (Homo sapiens)". Human Ageing Genomic Resources. Retrieved 2009-03-05.
- "Catalase". MadSci FAQ. madsci.org. Retrieved 2009-03-05.
- "Catalase and oxidase test video". Regnvm Prokaryotae. Retrieved 2009-03-05.
- "EC 1.11.1.6 - catalase". Brenda: The Comprehensive Enzyme Information System. Retrieved 2009-03-05.
- "PeroxiBase - The peroxidase database". Swiss Institute of Bioinformatics. Archived from the original on 2008-10-13. Retrieved 2009-03-05.
- "Catalase Procedure". MicrobeID.com. Retrieved 2009-04-22.
- "Catalase Molecule of the Month". Protein Data Bank. Archived from the original on 2013-05-11. Retrieved 2013-01-08.
- Overview of all the structural information available in the PDB for UniProt: P04040 (Catalase) at the PDBe-KB.